一种FnCpf1突变体的制作方法
本发明属于分子生物学领域,具体涉及一种fncpf1突变体。
背景技术:
二型限制限制性内切酶(re)的发现促进了dna重组技术的广泛应用。随后,合成生物学的飞速发展触发了对灵活多用的体外dna组装/编辑策略的需求。迄今,体外dna组装/编辑方法可分为两类:酶切连接和同源序列引导的同源重组。目前,最常用的酶切连接主要包括传统的基于限制性内切酶的酶切连接和goldengate组装。goldengate是基于iis的限制性内切酶的一步克隆方法,并已优化用于多片段dna的按序组装(werner,engleretal.2012)。但是,对于既定的dna序列,iis内切酶识别位点分布的不确定性和其存在数量的有限性,其编辑会受到限制。对于基于同源重组的体外编辑方法而言,例如gibson组装,片段数的增加或者片段中高gc序列的出现,编辑效率均会受到影响而下降。另外,由于其序列依赖性,我们无法实现对于高相似序列的重复编辑,例如将同一个启动子重复引入至目标基因的不同位置。这些编辑方法中存在的限制也强调了对于开发新方法的需求性和重要性。
快速增长的技术-规律成簇的间隔短回文重复(crispr)系统近几年发展迅速,其中最广泛应用的crispr系统是来自于化脓链球菌的crispr/cas9。该系统在研究者的设计和优化下,被用于辅助体外的分子克隆。但是,这些方法都是利用同源重组的方法将cas9剪切产生的平末端与目标载体相连,所以对于序列高相似性的dna片段的编辑往往会受到一定的局限。
2015年,张锋等发现并鉴定了一种新的第二类v型的系统crispr-cpf1,随后研究者发现并证明了其在多种有机体中的应用潜能和价值。相比于cas9的体外编辑,cpf1具有以下优势:1)cpf1剪切双链dna产生粘性末端;2)cpf1的引导不需要反向作用的rna,所以引导crrna的长度更短(~42bp);3)cpf1可以自己加工crrna簇形成单个成熟的crrna,使其易于开发用于多基因编辑。因此,基于cpf1的体外编辑方法可以为cas9的应用提供另一种选择性。
在众多已经鉴定的cpf1蛋白中,来源于francisellanovicida的fncpf1具有一个明显的优势-fncpf1识别的pam较短(3bp),并且当间歇序列的长度为23bp时,特异性的剪切pam远端的第18个碱基和互补链的第23碱基,产生5bp的粘性末端。由于cpf1的剪切由crrna(识别的间歇区通常为23bp)介导,相同的dna序列出现的概率大大减少,因此很好的规避了识别位点非理想带来的限制。综上,fncpf1介导的体外dna编辑方法可以解决很多现有技术上的缺陷,但是fncpf1识别范围限定于ttn,使用范围仍然有限。
技术实现要素:
为了解决前述问题,本发明提供了一种识别范围更广的cpf1突变体。
本发明fncpf1突变体,它是在野生型fncpf1基础上,经过氨基酸位点突变得到,所述野生型fncpf1的氨基酸序列如seqidno:1所示,所述突变的位点包含asn607、lys180、lys660、asp616中的至少一个突变。
其中,它包含asn607arg、lys180ser、lys660arg、asp616asn的至少一个突变。
asn607arg,是指野生型fncpf1第607位的氨基酸asn突变成为了arg,其余突变的解释方式相同。
其中,它还包含lys671arg突变、lys613val和/或asn617arg突变。
其中,它的氨基酸序列为seqidno:2或3所示。
本发明还提供了上述cpf1突变体的核苷酸序列。
本发明还提供了一种重组载体,它包含编码上述cpf1突变体的核苷酸序列;所述的重组载体是原核载体。
本发明还提供了一种重组菌,它包含上述的重组载体;优选地,所述细菌为大肠杆菌bl21(de3)菌株。
本发明还提供了制备上述cpf1突变体的方法。
本发明还提供了上述cpf1突变体、核苷酸序列、重组载体,或重组菌在基因编辑中的用途。
其中,所述基因编辑包含基因敲除、基因突变、靶向基因激活/抑制、dna连接、dna多片段组装、dna片段插入、dna片段替换、碱基替换。
seqidno:1:野生型fncpf1的氨基酸序列1-1300:msiyqefvnkyslsktlrfelipqgktlenikarglilddekrakdykkakqiidkyhqffieeilssvcisedllqnysdvyfklkksdddnlqkdfksakdtikkqiseyikdsekfknlfnqnlidakkgqesdlilwlkqskdngielfkansditdidealeiiksfkgwttyfkgfhenrknvyssndiptsiiyrivddnlpkflenkakyeslkdkapeainyeqikkdlaeeltfdidyktsevnqrvfsldevfeianfnnylnqsgitkfntiiggkfvngentkrkgineyinlysqqindktlkkykmsvlfkqilsdtesksfvidkleddsdvvttmqsfyeqiaafktveeksiketlsllfddlkaqkldlskiyfkndksltdlsqqvfddysvigtavleyitqqiapknldnpskkeqeliakktekakylsletiklaleefnkhrdidkqcrfeeilanfaaipmifdeiaqnkdnlaqisikyqnqgkkdllqasaeddvkaikdlldqtnnllhklkifhisqsedkanildkdehfylvfeecyfelanivplynkirnyitqkpysdekfklnfenstlangwdknkepdntailfikddkyylgvmnkknnkifddkaikenkgegykkivykllpgankmlpkvffsaksikfynpsedilrirnhsthtkngspqkgyekfefniedcrkfidfykqsiskhpewkdfgfrfsdtqrynsidefyrevenqgykltfenisesyidsvvnqgklylfqiynkdfsayskgrpnlhtlywkalfdernlqdvvyklngeaelfyrkqsipkkithpakeaianknkdnpkkesvfeydlikdkrftedkfffhcpitinfkssgankfndeinlllkekandvhilsidrgerhlayytlvdgkgniikqdtfniigndrmktnyhdklaaiekdrdsarkdwkkinnikemkegylsqvvheiaklvieynaivvfedlnfgfkrgrfkvekqvyqklekmlieklnylvfkdnefdktggvlrayqltapfetfkkmgkqtgiiyyvpagftskicpvtgfvnqlypkyesvsksqeffskfdkicynldkgyfefsfdyknfgdkaakgkwtiasfgsrlinfrnsdknhnwdtrevyptkelekllkdysieyghgecikaaicgesdkkffakltsvlntilqmrnsktgteldylispvadvngnffdsrqapknmpqdadangayhiglkglmllgriknnqegkklnlvikneeyfefvqnrnn*
seqidno:2:ep15的氨基酸序列
msiyqefvnkyslsktlrfelipqgktlenikarglilddekrakdykkakqiidkyhqffieeilssvcisedllqnysdvyfklkksdddnlqkdfksakdtikkqiseyikdsekfknlfnqnlidakkgqesdlilwlkqskdngielfkansditdidealeiiksfkgwttyfsgfhenrknvyssndiptsiiyrivddnlpkflenkakyeslkdkapeainyeqikkdlaeeltfdidyktsevnqrvfsldevfeianfnnylnqsgitkfntiiggkfvngentkrkgineyinlysqqindktlkkykmsvlfkqilsdtesksfvidkleddsdvvttmqsfyeqiaafktveeksiketlsllfddlkaqkldlskiyfkndksltdlsqqvfddysvigtavleyitqqiapknldnpskkeqeliakktekakylsletiklaleefnkhrdidkqcrfeeilanfaaipmifdeiaqnkdnlaqisikyqnqgkkdllqasaeddvkaikdlldqtnnllhklkifhisqsedkanildkdehfylvfeecyfelanivplynkirnyitqkpysdekfklnfenstlargwdknkepnntailfikddkyylgvmnkknnkifddkaikenkgegykkivyrllpgankmlprvffsaksikfynpsedilrirnhsthtkngspqkgyekfefniedcrkfidfykqsiskhpewkdfgfrfsdtqrynsidefyrevenqgykltfenisesyidsvvnqgklylfqiynkdfsayskgrpnlhtlywkalfdernlqdvvyklngeaelfyrkqsipkkithpakeaianknkdnpkkesvfeydlikdkrftedkfffhcpitinfkssgankfndeinlllkekandvhilsidrgerhlayytlvdgkgniikqdtfniigndrmktnyhdklaaiekdrdsarkdwkkinnikemkegylsqvvheiaklvieynaivvfedlnfgfkrgrfkvekqvyqklekmlieklnylvfkdnefdktggvlrayqltapfetfkkmgkqtgiiyyvpagftskicpvtgfvnqlypkyesvsksqeffskfdkicynldkgyfefsfdyknfgdkaakgkwtiasfgsrlinfrnsdknhnwdtrevyptkelekllkdysieyghgecikaaicgesdkkffakltsvlntilqmrnsktgteldylispvadvngnffdsrqapknmpqdadangayhiglkglmllgriknnqegkklnlvikneeyfefvqnrnn*
seqidno:3:ep16的氨基酸序列
msiyqefvnkyslsktlrfelipqgktlenikarglilddekrakdykkakqiidkyhqffieeilssvcisedllqnysdvyfklkksdddnlqkdfksakdtikkqiseyikdsekfknlfnqnlidakkgqesdlilwlkqskdngielfkansditdidealeiiksfkgwttyfsgfhenrknvyssndiptsiiyrivddnlpkflenkakyeslkdkapeainyeqikkdlaeeltfdidyktsevnqrvfsldevfeianfnnylnqsgitkfntiiggkfvngentkrkgineyinlysqqindktlkkykmsvlfkqilsdtesksfvidkleddsdvvttmqsfyeqiaafktveeksiketlsllfddlkaqkldlskiyfkndksltdlsqqvfddysvigtavleyitqqiapknldnpskkeqeliakktekakylsletiklaleefnkhrdidkqcrfeeilanfaaipmifdeiaqnkdnlaqisikyqnqgkkdllqasaeddvkaikdlldqtnnllhklkifhisqsedkanildkdehfylvfeecyfelanivplynkirnyitqkpysdekfklnfenstlargwdknvepnrtailfikddkyylgvmnkknnkifddkaikenkgegykkivyrllpgankmlpkvffsaksikfynpsedilrirnhsthtkngspqkgyekfefniedcrkfidfykqsiskhpewkdfgfrfsdtqrynsidefyrevenqgykltfenisesyidsvvnqgklylfqiynkdfsayskgrpnlhtlywkalfdernlqdvvyklngeaelfyrkqsipkkithpakeaianknkdnpkkesvfeydlikdkrftedkfffhcpitinfkssgankfndeinlllkekandvhilsidrgerhlayytlvdgkgniikqdtfniigndrmktnyhdklaaiekdrdsarkdwkkinnikemkegylsqvvheiaklvieynaivvfedlnfgfkrgrfkvekqvyqklekmlieklnylvfkdnefdktggvlrayqltapfetfkkmgkqtgiiyyvpagftskicpvtgfvnqlypkyesvsksqeffskfdkicynldkgyfefsfdyknfgdkaakgkwtiasfgsrlinfrnsdknhnwdtrevyptkelekllkdysieyghgecikaaicgesdkkffakltsvlntilqmrnsktgteldylispvadvngnffdsrqapknmpqdadangayhiglkglmllgriknnqegkklnlvikneeyfefvqnrnn*
ep15的核苷酸序列(seqidno:4)为:
atgagcatctatcaggagttcgtgaataagtacagcctgtccaagaccctgcggtttgagctgatcccccagggcaagacactggagaacatcaaggccaggggcctgatcctggacgatgagaagcgcgccaaggactataagaaggccaagcagatcatcgataagtaccaccagttctttatcgaggagatcctgagcagcgtgtgcatctctgaggatctgctgcagaattacagcgacgtgtatttcaagctgaagaagtctgacgatgacaacctgcagaaggacttcaagagcgccaaggacaccatcaagaagcagatcagcgagtatatcaaggactccgagaagtttaagaatctgttcaaccagaatctgatcgatgccaagaagggccaggagtccgacctgatcctgtggctgaagcagtctaaggacaatggcatcgagctgttcaaggccaactctgatatcaccgatatcgacgaggccctggagatcatcaagagctttaagggctggaccacatactttagcggcttccacgagaacaggaagaacgtgtacagcagcaacgacatccctacaagcatcatctaccgcatcgtggatgacaatctgccaaagttcctggagaacaaggccaagtatgagtccctgaaggacaaggcccccgaggccatcaattacgagcagatcaagaaggatctggccgaggagctgaccttcgatatcgactataagacatccgaggtgaaccagcgggtgttttctctggacgaggtgtttgagatcgccaatttcaacaattacctgaaccagtccggcatcaccaagttcaatacaatcatcggcggcaagtttgtgaacggcgagaataccaagagaaagggcatcaacgagtacatcaatctgtatagccagcagatcaacgacaagaccctgaagaagtacaagatgagcgtgctgttcaagcagatcctgtccgatacagagtctaagagctttgtgatcgataagctggaggatgactctgacgtggtgaccacaatgcagagcttttatgagcagatcgccgccttcaagaccgtggaggagaagtctatcaaggagacactgagcctgctgttcgatgacctgaaggcccagaagctggacctgtctaagatctacttcaagaacgataagtccctgaccgacctgtctcagcaggtgtttgatgactatagcgtgatcggcaccgccgtgctggagtacatcacacagcagatcgccccaaagaacctggataatccctctaagaaggagcaggagctgatcgccaagaagaccgagaaggccaagtatctgagcctggagacaatcaagctggccctggaggagttcaataagcaccgggatatcgacaagcagtgcagatttgaggagatcctggccaacttcgccgccatccccatgatctttgatgagatcgcccagaacaaggacaatctggcccagatctccatcaagtaccagaaccagggcaagaaggacctgctgcaggcctctgccgaggatgacgtgaaggccatcaaggatctgctggaccagaccaacaatctgctgcacaagctgaagatcttccacatctcccagtctgaggataaggccaatatcctggataaggacgagcacttttatctggtgttcgaggagtgttacttcgagctggccaacatcgtgcccctgtacaacaagatcagaaattatatcacacagaagccttactccgacgagaagtttaagctgaacttcgagaacagcaccctggccagaggctgggataagaataaggagcctaacaacacagccatcctgttcatcaaggatgacaagtactatctgggcgtgatgaataagaagaacaataagatcttcgatgacaaggccatcaaggagaacaagggcgagggctacaagaagatcgtgtataggctgctgcccggcgccaataagatgctgcctagggtgttcttttccgccaagtctatcaagttctacaacccatccgaggacatcctgcggatcagaaatcactccacccacacaaagaacggctctccccagaagggctatgagaagtttgagttcaatatcgaggattgccggaagtttatcgacttctacaagcagagcatctccaagcaccctgagtggaaggattttggcttcaggtttagcgacacccagcggtacaactccatcgacgagttctacagagaggtggagaatcagggctataagctgacatttgagaacatctctgagagctacatcgacagcgtggtgaatcagggcaagctgtacctgttccagatctataacaaggacttcagcgcctattccaagggccggccaaacctgcacaccctgtactggaaggccctgttcgatgagagaaatctgcaggacgtggtgtataagctgaacggcgaggccgagctgttttacaggaagcagtccatccctaagaagatcacacacccagccaaggaggccatcgccaacaagaataaggacaatcctaagaaggagagcgtgttcgagtacgatctgatcaaggacaagcggttcaccgaggataagttctttttccactgtccaatcacaatcaacttcaagtcctctggcgccaacaagtttaatgacgagatcaatctgctgctgaaggagaaggccaacgatgtgcacatcctgagcatcgaccggggcgagagacacctggcctactataccctggtggatggcaagggcaatatcatcaagcaggataccttcaacatcatcggcaatgacaggatgaagacaaactaccacgataagctggccgccatcgagaaggatagggactccgcccgcaaggactggaagaagatcaacaatatcaaggagatgaaggagggctatctgtctcaggtggtgcacgagatcgccaagctggtcatcgagtacaatgccatcgtggtgttcgaggatctgaacttcggctttaagaggggccgctttaaggtggagaagcaggtgtatcagaagctggagaagatgctgatcgagaagctgaattacctggtgtttaaggataacgagttcgacaagaccggaggcgtgctgagggcataccagctgaccgccccctttgagacattcaagaagatgggcaagcagacaggcatcatctactatgtgccagccggcttcacctccaagatctgccccgtgacaggctttgtgaaccagctgtaccctaagtatgagtccgtgtctaagagccaggagtttttcagcaagttcgataagatctgttataatctggacaagggctacttcgagttttccttcgattataagaactttggcgacaaggccgccaagggcaagtggaccatcgcctctttcggcagccggctgatcaactttagaaattccgataagaaccacaattgggacacccgggaggtgtacccaacaaaggagctggagaagctgctgaaggactacagcatcgagtatggccacggcgagtgcatcaaggccgccatctgtggcgagagcgataagaagtttttcgccaagctgacctccgtgctgaatacaatcctgcagatgcggaacagcaagaccggcacagagctggactacctgatctcccccgtggccgatgtgaacggcaacttcttcgacagcagacaggcccccaagaatatgcctcaggatgccgacgccaacggcgcctatcacatcggcctgaagggcctgatgctgctgggcaggatcaagaacaatcaggagggcaagaagctgaacctggtcatcaagaacgaggagtactttgagttcgtgcagaaccgcaacaattga
ep16的核苷酸序列(seqidno:5)为:
atgagcatctatcaggagttcgtgaataagtacagcctgtccaagaccctgcggtttgagctgatcccccagggcaagacactggagaacatcaaggccaggggcctgatcctggacgatgagaagcgcgccaaggactataagaaggccaagcagatcatcgataagtaccaccagttctttatcgaggagatcctgagcagcgtgtgcatctctgaggatctgctgcagaattacagcgacgtgtatttcaagctgaagaagtctgacgatgacaacctgcagaaggacttcaagagcgccaaggacaccatcaagaagcagatcagcgagtatatcaaggactccgagaagtttaagaatctgttcaaccagaatctgatcgatgccaagaagggccaggagtccgacctgatcctgtggctgaagcagtctaaggacaatggcatcgagctgttcaaggccaactctgatatcaccgatatcgacgaggccctggagatcatcaagagctttaagggctggaccacatactttagcggcttccacgagaacaggaagaacgtgtacagcagcaacgacatccctacaagcatcatctaccgcatcgtggatgacaatctgccaaagttcctggagaacaaggccaagtatgagtccctgaaggacaaggcccccgaggccatcaattacgagcagatcaagaaggatctggccgaggagctgaccttcgatatcgactataagacatccgaggtgaaccagcgggtgttttctctggacgaggtgtttgagatcgccaatttcaacaattacctgaaccagtccggcatcaccaagttcaatacaatcatcggcggcaagtttgtgaacggcgagaataccaagagaaagggcatcaacgagtacatcaatctgtatagccagcagatcaacgacaagaccctgaagaagtacaagatgagcgtgctgttcaagcagatcctgtccgatacagagtctaagagctttgtgatcgataagctggaggatgactctgacgtggtgaccacaatgcagagcttttatgagcagatcgccgccttcaagaccgtggaggagaagtctatcaaggagacactgagcctgctgttcgatgacctgaaggcccagaagctggacctgtctaagatctacttcaagaacgataagtccctgaccgacctgtctcagcaggtgtttgatgactatagcgtgatcggcaccgccgtgctggagtacatcacacagcagatcgccccaaagaacctggataatccctctaagaaggagcaggagctgatcgccaagaagaccgagaaggccaagtatctgagcctggagacaatcaagctggccctggaggagttcaataagcaccgggatatcgacaagcagtgcagatttgaggagatcctggccaacttcgccgccatccccatgatctttgatgagatcgcccagaacaaggacaatctggcccagatctccatcaagtaccagaaccagggcaagaaggacctgctgcaggcctctgccgaggatgacgtgaaggccatcaaggatctgctggaccagaccaacaatctgctgcacaagctgaagatcttccacatctcccagtctgaggataaggccaatatcctggataaggacgagcacttttatctggtgttcgaggagtgttacttcgagctggccaacatcgtgcccctgtacaacaagatcagaaattatatcacacagaagccttactccgacgagaagtttaagctgaacttcgagaacagcaccctggccagaggctgggataagaatgtggagcctaacagaacagccatcctgttcatcaaggatgacaagtactatctgggcgtgatgaataagaagaacaataagatcttcgatgacaaggccatcaaggagaacaagggcgagggctacaagaagatcgtgtataggctgctgcccggcgccaataagatgctgcctaaggtgttcttttccgccaagtctatcaagttctacaacccatccgaggacatcctgcggatcagaaatcactccacccacacaaagaacggctctccccagaagggctatgagaagtttgagttcaatatcgaggattgccggaagtttatcgacttctacaagcagagcatctccaagcaccctgagtggaaggattttggcttcaggtttagcgacacccagcggtacaactccatcgacgagttctacagagaggtggagaatcagggctataagctgacatttgagaacatctctgagagctacatcgacagcgtggtgaatcagggcaagctgtacctgttccagatctataacaaggacttcagcgcctattccaagggccggccaaacctgcacaccctgtactggaaggccctgttcgatgagagaaatctgcaggacgtggtgtataagctgaacggcgaggccgagctgttttacaggaagcagtccatccctaagaagatcacacacccagccaaggaggccatcgccaacaagaataaggacaatcctaagaaggagagcgtgttcgagtacgatctgatcaaggacaagcggttcaccgaggataagttctttttccactgtccaatcacaatcaacttcaagtcctctggcgccaacaagtttaatgacgagatcaatctgctgctgaaggagaaggccaacgatgtgcacatcctgagcatcgaccggggcgagagacacctggcctactataccctggtggatggcaagggcaatatcatcaagcaggataccttcaacatcatcggcaatgacaggatgaagacaaactaccacgataagctggccgccatcgagaaggatagggactccgcccgcaaggactggaagaagatcaacaatatcaaggagatgaaggagggctatctgtctcaggtggtgcacgagatcgccaagctggtcatcgagtacaatgccatcgtggtgttcgaggatctgaacttcggctttaagaggggccgctttaaggtggagaagcaggtgtatcagaagctggagaagatgctgatcgagaagctgaattacctggtgtttaaggataacgagttcgacaagaccggaggcgtgctgagggcataccagctgaccgccccctttgagacattcaagaagatgggcaagcagacaggcatcatctactatgtgccagccggcttcacctccaagatctgccccgtgacaggctttgtgaaccagctgtaccctaagtatgagtccgtgtctaagagccaggagtttttcagcaagttcgataagatctgttataatctggacaagggctacttcgagttttccttcgattataagaactttggcgacaaggccgccaagggcaagtggaccatcgcctctttcggcagccggctgatcaactttagaaattccgataagaaccacaattgggacacccgggaggtgtacccaacaaaggagctggagaagctgctgaaggactacagcatcgagtatggccacggcgagtgcatcaaggccgccatctgtggcgagagcgataagaagtttttcgccaagctgacctccgtgctgaatacaatcctgcagatgcggaacagcaagaccggcacagagctggactacctgatctcccccgtggccgatgtgaacggcaacttcttcgacagcagacaggcccccaagaatatgcctcaggatgccgacgccaacggcgcctatcacatcggcctgaagggcctgatgctgctgggcaggatcaagaacaatcaggagggcaagaagctgaacctggtcatcaagaacgaggagtactttgagttcgtgcagaaccgcaacaattga
本发明改进后的fncpf1突变体对序列的识别范围广,是野生型fncpf1识别范围的2~2.5倍,可以应用于crispr-cpf1系统,编辑dna片段,可以克服传统野生型fncpf1识别范围太窄,应用范围太窄的问题,应用前景优良。
显然,根据本发明的上述内容,按照本领域的普通技术知识和惯用手段,在不脱离本发明上述基本技术思想前提下,还可以做出其它多种形式的修改、替换或变更。
以下通过实施例形式的具体实施方式,对本发明的上述内容再作进一步的详细说明。但不应将此理解为本发明上述主题的范围仅限于以下的实例。凡基于本发明上述内容所实现的技术均属于本发明的范围。
附图说明
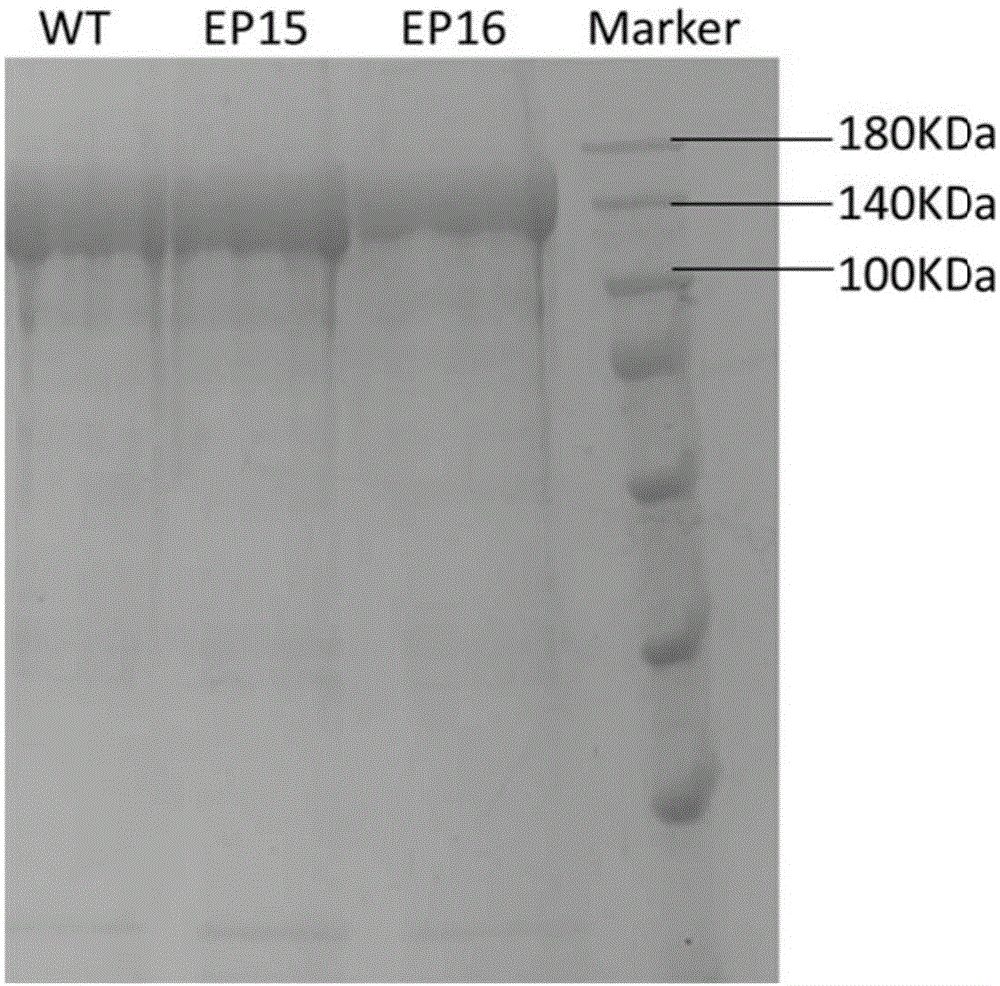
图1野生型fncpf1及突变体蛋白ep15和ep16的sds-page电泳图谱,cpf1蛋白大小151kda
图2野生fncpf1的pam检测结果图,模板大小:1234bp,产物大小:707bp和527bp;
图3fncpf1突变体ep15的pam检测结果图,模板大小:1234bp,产物大小:707bp和527bp;
图4fncpf1突变体ep16的pam检测结果图,模板大小:1234bp,产物大小:707bp和527bp。
具体实施方式
下面以实施例作进一步说明,但本发明不局限于这些实施例。
实施例1fncpf1突变体的制备以及效果验证一、酶的制备(克隆:表达和纯化野生型和突变型fncpf1蛋白)
1、制备突变质粒
1)fncpf1(wp003040289)表达质粒来源于上海中科院。随后fncpf1突变体的表达质粒以此为基础,进行定点突变pcr,在:野生型fncpf1(氨基酸序列如seqidno:1所示)的基础上,拟突变制备突变体ep15和ep16,其中ep15的突变为:lys180ser、asn607arg、asp616asn、lys660arg和lys671arg突变,ep16的突变为:lys180ser、asn607arg、lys613val、asp616asn、asn617arg和lys660arg。
2)根据拟突变的氨基酸序列进行设计引物,引物由北京擎科新业生物技术有限公司合成,所用引物见表1:
表1用于获得突变蛋白的引物
利用所得的引物进行定点突变pcr,模板为pet28tev-fncpf1质粒,所用酶为q5高保真酶(neb公司,货号m0491)。具体配置体系如下:
反应程序为:98℃,2min;反应98℃,10s;退火,10s;60℃,5min;72℃,5min;4℃,保存;运行25个循环。
对pcr产物进行胶回收(北京天根生化科技有限公司,货号dp214),随后利用限制性内切酶dpni(neb公司,货号r0176l)去除模板(37℃,2h),将上述所得反应液转化大肠杆菌感受态细胞(dh5α)中,涂布于lb(含50μg/ml卡那抗生素)平板中,置于37℃培养箱中过夜。挑取平板中的单克隆,测序(北京擎科新业生物技术有限公司),获得含有目标突变(目标片段如)的正确克隆。
2、野生型及突变体蛋白的表达纯化
蛋白纯化的步骤为:
1)将表达质粒转化至e.colibl21(de3);
2)挑单克隆加到3ml含有50ug/mlkanamycin的lb液体培养基中37℃,220rpm过夜培养16小时;
3)将过夜培养菌液按1:1000加到含有50ug/mlkanamycin的lb液体培养基中,在37℃,200rpm培养至od600=0.2左右,随后将培养箱降温至20℃,继续220rpm培养直到od600达到0.6-0.8;
4)加入iptg诱导剂,随后进行20℃,220rpm过夜培养;
5)将过夜菌液4℃,3800rpm离心10分钟收菌,弃上清,将装有菌块的离心瓶置于冰上,用30ml含有1mmdtt和1mmpmsf的预冷1xcpf1lysisbuffer重悬菌块;
6)将重悬后的菌液转入50ml国产bd管中,置于冰上,随后使用sonicator进行超声破菌(参数设置为30%强度,开3秒,停6秒,超声15分钟);
7)将细菌裂解液转入高速塑料离心管中,进行40c,18000rpm,30分钟高速离心;
8)高速离心期间,用5-10cv预冷的1xcpf1washingbuffer对镍柱进行平衡;
9)将7)中的上清液转入50mlbd管中,并加入8)中平衡过的镍胶与其混合,在层析柜中旋转混合一个小时使his-tag与镍结合;
10)将9中的蛋白镍胶混合液重新转入空柱中,用250ml预冷的1xcpf1washingbuffer洗掉未与镍胶结合的杂蛋白,收集第一批穿出样本用于跑胶的阴性对照;
11)随后加入20ml预冷的1xcpf1elutionbuffer洗脱目标蛋白,保留少量洗脱样本用于跑胶;
12)使用nanodrop测量蛋白浓度,然后按照1mg的tev酶去切100mg的目标蛋白的比例加tev酶进行过夜剪切;
13)使用millipore离心浓缩管将tev酶过夜酶切的目标蛋白样本浓缩至0.2-0.5ml,随后加入50ml预冷的1xcpf1washingbuffer稀释样本中的咪唑浓度;
14)样本浓缩期间,用5-10cv预冷的1xcpf1washingbuffer对镍柱进行平衡,为反挂镍做准备;
15)将13)和14)混合,按照9进行操作;
16)将15)中的样本重新转入空柱中,收集含有切掉his-tag的目标蛋白的穿出液,随后用1xelutionbuffer洗脱镍胶,洗脱液做为tev酶的酶切效果的阴性对照;
17)测量16)中收集的穿出液,按照浓度进行进一步的浓缩;
18)进行12%sdg-page检测蛋白纯度;
19)制备50%(v/v)甘油cpf1蛋白储存样本,速冻并保存在-80℃中。
采用上述方法,制备得到野生型fncpf1及突变体蛋白p15和p16,采用电泳等方式验证的确得到了野生型fncpf1及突变体蛋白p15和p16(如图1)。
二、酶活的验证
由于fncpf1识别的pam为3位,故设计含有pam为5’-nnn-spacer-3’的dna底物,用于酶活的验证。根据反应需要,合成与底物对应的crrna。
1、crrna合成
1)转录的模板dna制备:根据crrna需要,设计合成转录的模板dna对应的引物如下表2:
表2用于pam测试的crrna制备引物
利用所得的引物进行dna富集pcr,具体配置体系如下:
反应程序为:98℃,2min;反应98℃,10s;退火,10s;50℃,7s;72℃,5min;4℃,保存;运行35个循环。
pcr反应液采用dna纯化试剂盒回收(tiangen,beijing,china)。
1)crrna的合成:rna的合成参照hiscribetmt7highyieldrnasynthesiskit(neb)的标准操作方案。制备好的rna采用rnaclean&concentrator-5kit(zymoresearch,ca,usa)回收。浓度测量采用nanodrop2000c(thermofisherscientific,massachusetts,usa)。rna稀释至20μm备用;
2)底物dna样品制备
为了制备具有对比性的dna模板,我们首先制备了一系列包含同一spacer不同pam的质粒库(表3)。然后利用pcr的方式富集得到64种pam不同的1300bp的dna底物;pcr引物如(表4)
表3pam验证质粒库
表4pam测试dna底物扩增引物
利用所得的引物进行dna富集pcr,具体配置体系如下:
反应程序为:98℃,2min;反应98℃,10s;退火,10s;50℃,7s;72℃,5min;4℃,保存;运行35个循环。
反应液采用dna纯化试剂盒回收(tiangen,beijing,china),浓度测量采用nanodrop2000c(thermofisherscientific,massachusetts,usa)。模板dna(对应表3标记为1-64)稀释至100ng/μl备用;
2、酶切反应及检测
表5反应加样
反应条件:37℃反应3h,然后75℃处理10min失活;然后向反应液加入适宜比例的loadingdye,neb,(6x);
凝胶成像采用geldoctmxr+withimagelabtmsoftware(bio-rad,california,usa),并通过对应软件分析各条带的体积值,通过公式计算得到pam体外效率。
[e=100x(((1-(1-(b+c)/(a+b+c))^1/2)]。
3、结果
凝胶成像结果如图2~图4所示,对应分析得到剪切效率统计为表6:
表6pam鉴定结果
由表6可以看出,野生型fncpf1对tta等9种序列的识别效率高,而经过本发明改进后的fncpf1突变体ep15对tta等18种序列的识别效果率均非常高,fncpf1突变体ep16对24种序列的识别效果率均非常高。
实验结果说明,本发明改进后的突变体对序列的识别范围广,可以用于更多序列的编辑,应用范围更广,实际应用前景更优良。
sequencelisting
<110>四川大学华西医院
<120>一种fncpf1突变体
<130>gy026-2019p013441cc
<150>201810393544.7
<151>2018-04-27
<160>5
<170>patentinversion3.5
<210>1
<211>1300
<212>prt
<213>francisellanovicida(野生型fncpf1的氨基酸序)
<400>1
metseriletyrglngluphevalasnlystyrserleuserlysthr
151015
leuargphegluleuileproglnglylysthrleugluasnilelys
202530
alaargglyleuileleuaspaspglulysargalalysasptyrlys
354045
lysalalysglnileileasplystyrhisglnphepheilegluglu
505560
ileleuserservalcysilesergluaspleuleuglnasntyrser
65707580
aspvaltyrphelysleulyslysseraspaspaspasnleuglnlys
859095
aspphelysseralalysaspthrilelyslysglnileserglutyr
100105110
ilelysaspserglulysphelysasnleupheasnglnasnleuile
115120125
aspalalyslysglyglngluseraspleuileleutrpleulysgln
130135140
serlysaspasnglyilegluleuphelysalaasnseraspilethr
145150155160
aspileaspglualaleugluileilelysserphelysglytrpthr
165170175
thrtyrphelysglyphehisgluasnarglysasnvaltyrserser
180185190
asnaspileprothrserileiletyrargilevalaspaspasnleu
195200205
prolyspheleugluasnlysalalystyrgluserleulysasplys
210215220
alaproglualaileasntyrgluglnilelyslysaspleualaglu
225230235240
gluleuthrpheaspileasptyrlysthrsergluvalasnglnarg
245250255
valpheserleuaspgluvalphegluilealaasnpheasnasntyr
260265270
leuasnglnserglyilethrlyspheasnthrileileglyglylys
275280285
phevalasnglygluasnthrlysarglysglyileasnglutyrile
290295300
asnleutyrserglnglnileasnasplysthrleulyslystyrlys
305310315320
metservalleuphelysglnileleuseraspthrgluserlysser
325330335
phevalileasplysleugluaspaspseraspvalvalthrthrmet
340345350
glnserphetyrgluglnilealaalaphelysthrvalgluglulys
355360365
serilelysgluthrleuserleuleupheaspaspleulysalagln
370375380
lysleuaspleuserlysiletyrphelysasnasplysserleuthr
385390395400
aspleuserglnglnvalpheaspasptyrservalileglythrala
405410415
valleuglutyrilethrglnglnilealaprolysasnleuaspasn
420425430
proserlyslysgluglngluleuilealalyslysthrglulysala
435440445
lystyrleuserleugluthrilelysleualaleugluglupheasn
450455460
lyshisargaspileasplysglncysargpheglugluileleuala
465470475480
asnphealaalaileprometilepheaspgluilealaglnasnlys
485490495
aspasnleualaglnileserilelystyrglnasnglnglylyslys
500505510
aspleuleuglnalaseralagluaspaspvallysalailelysasp
515520525
leuleuaspglnthrasnasnleuleuhislysleulysilephehis
530535540
ileserglnsergluasplysalaasnileleuasplysaspgluhis
545550555560
phetyrleuvalphegluglucystyrphegluleualaasnileval
565570575
proleutyrasnlysileargasntyrilethrglnlysprotyrser
580585590
aspglulysphelysleuasnphegluasnserthrleualaasngly
595600605
trpasplysasnlysgluproaspasnthralaileleupheilelys
610615620
aspasplystyrtyrleuglyvalmetasnlyslysasnasnlysile
625630635640
pheaspasplysalailelysgluasnlysglygluglytyrlyslys
645650655
ilevaltyrlysleuleuproglyalaasnlysmetleuprolysval
660665670
phepheseralalysserilelysphetyrasnprosergluaspile
675680685
leuargileargasnhisserthrhisthrlysasnglyserprogln
690695700
lysglytyrglulyspheglupheasnilegluaspcysarglysphe
705710715720
ileaspphetyrlysglnserileserlyshisproglutrplysasp
725730735
pheglypheargpheseraspthrglnargtyrasnserileaspglu
740745750
phetyrarggluvalgluasnglnglytyrlysleuthrphegluasn
755760765
ileserglusertyrileaspservalvalasnglnglylysleutyr
770775780
leupheglniletyrasnlysasppheseralatyrserlysglyarg
785790795800
proasnleuhisthrleutyrtrplysalaleupheaspgluargasn
805810815
leuglnaspvalvaltyrlysleuasnglyglualagluleuphetyr
820825830
arglysglnserileprolyslysilethrhisproalalysgluala
835840845
ilealaasnlysasnlysaspasnprolyslysgluservalpheglu
850855860
tyraspleuilelysasplysargphethrgluasplysphephephe
865870875880
hiscysproilethrileasnphelysserserglyalaasnlysphe
885890895
asnaspgluileasnleuleuleulysglulysalaasnaspvalhis
900905910
ileleuserileaspargglygluarghisleualatyrtyrthrleu
915920925
valaspglylysglyasnileilelysglnaspthrpheasnileile
930935940
glyasnaspargmetlysthrasntyrhisasplysleualaalaile
945950955960
glulysaspargaspseralaarglysasptrplyslysileasnasn
965970975
ilelysglumetlysgluglytyrleuserglnvalvalhisgluile
980985990
alalysleuvalileglutyrasnalailevalvalphegluaspleu
99510001005
asnpheglyphelysargglyargphelysvalglulysglnval
101010151020
tyrglnlysleuglulysmetleuileglulysleuasntyrleu
102510301035
valphelysaspasnglupheasplysthrglyglyvalleuarg
104010451050
alatyrglnleuthralaprophegluthrphelyslysmetgly
105510601065
lysglnthrglyileiletyrtyrvalproalaglyphethrser
107010751080
lysilecysprovalthrglyphevalasnglnleutyrprolys
108510901095
tyrgluservalserlysserglngluphepheserlyspheasp
110011051110
lysilecystyrasnleuasplysglytyrpheglupheserphe
111511201125
asptyrlysasnpheglyasplysalaalalysglylystrpthr
113011351140
ilealaserpheglyserargleuileasnpheargasnserasp
114511501155
lysasnhisasntrpaspthrarggluvaltyrprothrlysglu
116011651170
leuglulysleuleulysasptyrserileglutyrglyhisgly
117511801185
glucysilelysalaalailecysglygluserasplyslysphe
119011951200
phealalysleuthrservalleuasnthrileleuglnmetarg
120512101215
asnserlysthrglythrgluleuasptyrleuileserproval
122012251230
alaaspvalasnglyasnphepheaspserargglnalaprolys
123512401245
asnmetproglnaspalaaspalaasnglyalatyrhisilegly
125012551260
leulysglyleumetleuleuglyargilelysasnasnglnglu
126512701275
glylyslysleuasnleuvalilelysasngluglutyrpheglu
128012851290
phevalglnasnargasnasn
12951300
<210>2
<211>1300
<212>prt
<213>artificialsequence
<220>
<223>ep15的氨基酸序列
<400>2
metseriletyrglngluphevalasnlystyrserleuserlysthr
151015
leuargphegluleuileproglnglylysthrleugluasnilelys
202530
alaargglyleuileleuaspaspglulysargalalysasptyrlys
354045
lysalalysglnileileasplystyrhisglnphepheilegluglu
505560
ileleuserservalcysilesergluaspleuleuglnasntyrser
65707580
aspvaltyrphelysleulyslysseraspaspaspasnleuglnlys
859095
aspphelysseralalysaspthrilelyslysglnileserglutyr
100105110
ilelysaspserglulysphelysasnleupheasnglnasnleuile
115120125
aspalalyslysglyglngluseraspleuileleutrpleulysgln
130135140
serlysaspasnglyilegluleuphelysalaasnseraspilethr
145150155160
aspileaspglualaleugluileilelysserphelysglytrpthr
165170175
thrtyrpheserglyphehisgluasnarglysasnvaltyrserser
180185190
asnaspileprothrserileiletyrargilevalaspaspasnleu
195200205
prolyspheleugluasnlysalalystyrgluserleulysasplys
210215220
alaproglualaileasntyrgluglnilelyslysaspleualaglu
225230235240
gluleuthrpheaspileasptyrlysthrsergluvalasnglnarg
245250255
valpheserleuaspgluvalphegluilealaasnpheasnasntyr
260265270
leuasnglnserglyilethrlyspheasnthrileileglyglylys
275280285
phevalasnglygluasnthrlysarglysglyileasnglutyrile
290295300
asnleutyrserglnglnileasnasplysthrleulyslystyrlys
305310315320
metservalleuphelysglnileleuseraspthrgluserlysser
325330335
phevalileasplysleugluaspaspseraspvalvalthrthrmet
340345350
glnserphetyrgluglnilealaalaphelysthrvalgluglulys
355360365
serilelysgluthrleuserleuleupheaspaspleulysalagln
370375380
lysleuaspleuserlysiletyrphelysasnasplysserleuthr
385390395400
aspleuserglnglnvalpheaspasptyrservalileglythrala
405410415
valleuglutyrilethrglnglnilealaprolysasnleuaspasn
420425430
proserlyslysgluglngluleuilealalyslysthrglulysala
435440445
lystyrleuserleugluthrilelysleualaleugluglupheasn
450455460
lyshisargaspileasplysglncysargpheglugluileleuala
465470475480
asnphealaalaileprometilepheaspgluilealaglnasnlys
485490495
aspasnleualaglnileserilelystyrglnasnglnglylyslys
500505510
aspleuleuglnalaseralagluaspaspvallysalailelysasp
515520525
leuleuaspglnthrasnasnleuleuhislysleulysilephehis
530535540
ileserglnsergluasplysalaasnileleuasplysaspgluhis
545550555560
phetyrleuvalphegluglucystyrphegluleualaasnileval
565570575
proleutyrasnlysileargasntyrilethrglnlysprotyrser
580585590
aspglulysphelysleuasnphegluasnserthrleualaarggly
595600605
trpasplysasnlysgluproasnasnthralaileleupheilelys
610615620
aspasplystyrtyrleuglyvalmetasnlyslysasnasnlysile
625630635640
pheaspasplysalailelysgluasnlysglygluglytyrlyslys
645650655
ilevaltyrargleuleuproglyalaasnlysmetleuproargval
660665670
phepheseralalysserilelysphetyrasnprosergluaspile
675680685
leuargileargasnhisserthrhisthrlysasnglyserprogln
690695700
lysglytyrglulyspheglupheasnilegluaspcysarglysphe
705710715720
ileaspphetyrlysglnserileserlyshisproglutrplysasp
725730735
pheglypheargpheseraspthrglnargtyrasnserileaspglu
740745750
phetyrarggluvalgluasnglnglytyrlysleuthrphegluasn
755760765
ileserglusertyrileaspservalvalasnglnglylysleutyr
770775780
leupheglniletyrasnlysasppheseralatyrserlysglyarg
785790795800
proasnleuhisthrleutyrtrplysalaleupheaspgluargasn
805810815
leuglnaspvalvaltyrlysleuasnglyglualagluleuphetyr
820825830
arglysglnserileprolyslysilethrhisproalalysgluala
835840845
ilealaasnlysasnlysaspasnprolyslysgluservalpheglu
850855860
tyraspleuilelysasplysargphethrgluasplysphephephe
865870875880
hiscysproilethrileasnphelysserserglyalaasnlysphe
885890895
asnaspgluileasnleuleuleulysglulysalaasnaspvalhis
900905910
ileleuserileaspargglygluarghisleualatyrtyrthrleu
915920925
valaspglylysglyasnileilelysglnaspthrpheasnileile
930935940
glyasnaspargmetlysthrasntyrhisasplysleualaalaile
945950955960
glulysaspargaspseralaarglysasptrplyslysileasnasn
965970975
ilelysglumetlysgluglytyrleuserglnvalvalhisgluile
980985990
alalysleuvalileglutyrasnalailevalvalphegluaspleu
99510001005
asnpheglyphelysargglyargphelysvalglulysglnval
101010151020
tyrglnlysleuglulysmetleuileglulysleuasntyrleu
102510301035
valphelysaspasnglupheasplysthrglyglyvalleuarg
104010451050
alatyrglnleuthralaprophegluthrphelyslysmetgly
105510601065
lysglnthrglyileiletyrtyrvalproalaglyphethrser
107010751080
lysilecysprovalthrglyphevalasnglnleutyrprolys
108510901095
tyrgluservalserlysserglngluphepheserlyspheasp
110011051110
lysilecystyrasnleuasplysglytyrpheglupheserphe
111511201125
asptyrlysasnpheglyasplysalaalalysglylystrpthr
113011351140
ilealaserpheglyserargleuileasnpheargasnserasp
114511501155
lysasnhisasntrpaspthrarggluvaltyrprothrlysglu
116011651170
leuglulysleuleulysasptyrserileglutyrglyhisgly
117511801185
glucysilelysalaalailecysglygluserasplyslysphe
119011951200
phealalysleuthrservalleuasnthrileleuglnmetarg
120512101215
asnserlysthrglythrgluleuasptyrleuileserproval
122012251230
alaaspvalasnglyasnphepheaspserargglnalaprolys
123512401245
asnmetproglnaspalaaspalaasnglyalatyrhisilegly
125012551260
leulysglyleumetleuleuglyargilelysasnasnglnglu
126512701275
glylyslysleuasnleuvalilelysasngluglutyrpheglu
128012851290
phevalglnasnargasnasn
12951300
<210>3
<211>1300
<212>prt
<213>artificialsequence
<220>
<223>ep16的氨基酸序列
<400>3
metseriletyrglngluphevalasnlystyrserleuserlysthr
151015
leuargphegluleuileproglnglylysthrleugluasnilelys
202530
alaargglyleuileleuaspaspglulysargalalysasptyrlys
354045
lysalalysglnileileasplystyrhisglnphepheilegluglu
505560
ileleuserservalcysilesergluaspleuleuglnasntyrser
65707580
aspvaltyrphelysleulyslysseraspaspaspasnleuglnlys
859095
aspphelysseralalysaspthrilelyslysglnileserglutyr
100105110
ilelysaspserglulysphelysasnleupheasnglnasnleuile
115120125
aspalalyslysglyglngluseraspleuileleutrpleulysgln
130135140
serlysaspasnglyilegluleuphelysalaasnseraspilethr
145150155160
aspileaspglualaleugluileilelysserphelysglytrpthr
165170175
thrtyrpheserglyphehisgluasnarglysasnvaltyrserser
180185190
asnaspileprothrserileiletyrargilevalaspaspasnleu
195200205
prolyspheleugluasnlysalalystyrgluserleulysasplys
210215220
alaproglualaileasntyrgluglnilelyslysaspleualaglu
225230235240
gluleuthrpheaspileasptyrlysthrsergluvalasnglnarg
245250255
valpheserleuaspgluvalphegluilealaasnpheasnasntyr
260265270
leuasnglnserglyilethrlyspheasnthrileileglyglylys
275280285
phevalasnglygluasnthrlysarglysglyileasnglutyrile
290295300
asnleutyrserglnglnileasnasplysthrleulyslystyrlys
305310315320
metservalleuphelysglnileleuseraspthrgluserlysser
325330335
phevalileasplysleugluaspaspseraspvalvalthrthrmet
340345350
glnserphetyrgluglnilealaalaphelysthrvalgluglulys
355360365
serilelysgluthrleuserleuleupheaspaspleulysalagln
370375380
lysleuaspleuserlysiletyrphelysasnasplysserleuthr
385390395400
aspleuserglnglnvalpheaspasptyrservalileglythrala
405410415
valleuglutyrilethrglnglnilealaprolysasnleuaspasn
420425430
proserlyslysgluglngluleuilealalyslysthrglulysala
435440445
lystyrleuserleugluthrilelysleualaleugluglupheasn
450455460
lyshisargaspileasplysglncysargpheglugluileleuala
465470475480
asnphealaalaileprometilepheaspgluilealaglnasnlys
485490495
aspasnleualaglnileserilelystyrglnasnglnglylyslys
500505510
aspleuleuglnalaseralagluaspaspvallysalailelysasp
515520525
leuleuaspglnthrasnasnleuleuhislysleulysilephehis
530535540
ileserglnsergluasplysalaasnileleuasplysaspgluhis
545550555560
phetyrleuvalphegluglucystyrphegluleualaasnileval
565570575
proleutyrasnlysileargasntyrilethrglnlysprotyrser
580585590
aspglulysphelysleuasnphegluasnserthrleualaarggly
595600605
trpasplysasnvalgluproasnargthralaileleupheilelys
610615620
aspasplystyrtyrleuglyvalmetasnlyslysasnasnlysile
625630635640
pheaspasplysalailelysgluasnlysglygluglytyrlyslys
645650655
ilevaltyrargleuleuproglyalaasnlysmetleuprolysval
660665670
phepheseralalysserilelysphetyrasnprosergluaspile
675680685
leuargileargasnhisserthrhisthrlysasnglyserprogln
690695700
lysglytyrglulyspheglupheasnilegluaspcysarglysphe
705710715720
ileaspphetyrlysglnserileserlyshisproglutrplysasp
725730735
pheglypheargpheseraspthrglnargtyrasnserileaspglu
740745750
phetyrarggluvalgluasnglnglytyrlysleuthrphegluasn
755760765
ileserglusertyrileaspservalvalasnglnglylysleutyr
770775780
leupheglniletyrasnlysasppheseralatyrserlysglyarg
785790795800
proasnleuhisthrleutyrtrplysalaleupheaspgluargasn
805810815
leuglnaspvalvaltyrlysleuasnglyglualagluleuphetyr
820825830
arglysglnserileprolyslysilethrhisproalalysgluala
835840845
ilealaasnlysasnlysaspasnprolyslysgluservalpheglu
850855860
tyraspleuilelysasplysargphethrgluasplysphephephe
865870875880
hiscysproilethrileasnphelysserserglyalaasnlysphe
885890895
asnaspgluileasnleuleuleulysglulysalaasnaspvalhis
900905910
ileleuserileaspargglygluarghisleualatyrtyrthrleu
915920925
valaspglylysglyasnileilelysglnaspthrpheasnileile
930935940
glyasnaspargmetlysthrasntyrhisasplysleualaalaile
945950955960
glulysaspargaspseralaarglysasptrplyslysileasnasn
965970975
ilelysglumetlysgluglytyrleuserglnvalvalhisgluile
980985990
alalysleuvalileglutyrasnalailevalvalphegluaspleu
99510001005
asnpheglyphelysargglyargphelysvalglulysglnval
101010151020
tyrglnlysleuglulysmetleuileglulysleuasntyrleu
102510301035
valphelysaspasnglupheasplysthrglyglyvalleuarg
104010451050
alatyrglnleuthralaprophegluthrphelyslysmetgly
105510601065
lysglnthrglyileiletyrtyrvalproalaglyphethrser
107010751080
lysilecysprovalthrglyphevalasnglnleutyrprolys
108510901095
tyrgluservalserlysserglngluphepheserlyspheasp
110011051110
lysilecystyrasnleuasplysglytyrpheglupheserphe
111511201125
asptyrlysasnpheglyasplysalaalalysglylystrpthr
113011351140
ilealaserpheglyserargleuileasnpheargasnserasp
114511501155
lysasnhisasntrpaspthrarggluvaltyrprothrlysglu
116011651170
leuglulysleuleulysasptyrserileglutyrglyhisgly
117511801185
glucysilelysalaalailecysglygluserasplyslysphe
119011951200
phealalysleuthrservalleuasnthrileleuglnmetarg
120512101215
asnserlysthrglythrgluleuasptyrleuileserproval
122012251230
alaaspvalasnglyasnphepheaspserargglnalaprolys
123512401245
asnmetproglnaspalaaspalaasnglyalatyrhisilegly
125012551260
leulysglyleumetleuleuglyargilelysasnasnglnglu
126512701275
glylyslysleuasnleuvalilelysasngluglutyrpheglu
128012851290
phevalglnasnargasnasn
12951300
<210>4
<211>3903
<212>dna
<213>artificialsequence
<220>
<223>ep15的核苷酸序列
<400>4
atgagcatctatcaggagttcgtgaataagtacagcctgtccaagaccctgcggtttgag60
ctgatcccccagggcaagacactggagaacatcaaggccaggggcctgatcctggacgat120
gagaagcgcgccaaggactataagaaggccaagcagatcatcgataagtaccaccagttc180
tttatcgaggagatcctgagcagcgtgtgcatctctgaggatctgctgcagaattacagc240
gacgtgtatttcaagctgaagaagtctgacgatgacaacctgcagaaggacttcaagagc300
gccaaggacaccatcaagaagcagatcagcgagtatatcaaggactccgagaagtttaag360
aatctgttcaaccagaatctgatcgatgccaagaagggccaggagtccgacctgatcctg420
tggctgaagcagtctaaggacaatggcatcgagctgttcaaggccaactctgatatcacc480
gatatcgacgaggccctggagatcatcaagagctttaagggctggaccacatactttagc540
ggcttccacgagaacaggaagaacgtgtacagcagcaacgacatccctacaagcatcatc600
taccgcatcgtggatgacaatctgccaaagttcctggagaacaaggccaagtatgagtcc660
ctgaaggacaaggcccccgaggccatcaattacgagcagatcaagaaggatctggccgag720
gagctgaccttcgatatcgactataagacatccgaggtgaaccagcgggtgttttctctg780
gacgaggtgtttgagatcgccaatttcaacaattacctgaaccagtccggcatcaccaag840
ttcaatacaatcatcggcggcaagtttgtgaacggcgagaataccaagagaaagggcatc900
aacgagtacatcaatctgtatagccagcagatcaacgacaagaccctgaagaagtacaag960
atgagcgtgctgttcaagcagatcctgtccgatacagagtctaagagctttgtgatcgat1020
aagctggaggatgactctgacgtggtgaccacaatgcagagcttttatgagcagatcgcc1080
gccttcaagaccgtggaggagaagtctatcaaggagacactgagcctgctgttcgatgac1140
ctgaaggcccagaagctggacctgtctaagatctacttcaagaacgataagtccctgacc1200
gacctgtctcagcaggtgtttgatgactatagcgtgatcggcaccgccgtgctggagtac1260
atcacacagcagatcgccccaaagaacctggataatccctctaagaaggagcaggagctg1320
atcgccaagaagaccgagaaggccaagtatctgagcctggagacaatcaagctggccctg1380
gaggagttcaataagcaccgggatatcgacaagcagtgcagatttgaggagatcctggcc1440
aacttcgccgccatccccatgatctttgatgagatcgcccagaacaaggacaatctggcc1500
cagatctccatcaagtaccagaaccagggcaagaaggacctgctgcaggcctctgccgag1560
gatgacgtgaaggccatcaaggatctgctggaccagaccaacaatctgctgcacaagctg1620
aagatcttccacatctcccagtctgaggataaggccaatatcctggataaggacgagcac1680
ttttatctggtgttcgaggagtgttacttcgagctggccaacatcgtgcccctgtacaac1740
aagatcagaaattatatcacacagaagccttactccgacgagaagtttaagctgaacttc1800
gagaacagcaccctggccagaggctgggataagaataaggagcctaacaacacagccatc1860
ctgttcatcaaggatgacaagtactatctgggcgtgatgaataagaagaacaataagatc1920
ttcgatgacaaggccatcaaggagaacaagggcgagggctacaagaagatcgtgtatagg1980
ctgctgcccggcgccaataagatgctgcctagggtgttcttttccgccaagtctatcaag2040
ttctacaacccatccgaggacatcctgcggatcagaaatcactccacccacacaaagaac2100
ggctctccccagaagggctatgagaagtttgagttcaatatcgaggattgccggaagttt2160
atcgacttctacaagcagagcatctccaagcaccctgagtggaaggattttggcttcagg2220
tttagcgacacccagcggtacaactccatcgacgagttctacagagaggtggagaatcag2280
ggctataagctgacatttgagaacatctctgagagctacatcgacagcgtggtgaatcag2340
ggcaagctgtacctgttccagatctataacaaggacttcagcgcctattccaagggccgg2400
ccaaacctgcacaccctgtactggaaggccctgttcgatgagagaaatctgcaggacgtg2460
gtgtataagctgaacggcgaggccgagctgttttacaggaagcagtccatccctaagaag2520
atcacacacccagccaaggaggccatcgccaacaagaataaggacaatcctaagaaggag2580
agcgtgttcgagtacgatctgatcaaggacaagcggttcaccgaggataagttctttttc2640
cactgtccaatcacaatcaacttcaagtcctctggcgccaacaagtttaatgacgagatc2700
aatctgctgctgaaggagaaggccaacgatgtgcacatcctgagcatcgaccggggcgag2760
agacacctggcctactataccctggtggatggcaagggcaatatcatcaagcaggatacc2820
ttcaacatcatcggcaatgacaggatgaagacaaactaccacgataagctggccgccatc2880
gagaaggatagggactccgcccgcaaggactggaagaagatcaacaatatcaaggagatg2940
aaggagggctatctgtctcaggtggtgcacgagatcgccaagctggtcatcgagtacaat3000
gccatcgtggtgttcgaggatctgaacttcggctttaagaggggccgctttaaggtggag3060
aagcaggtgtatcagaagctggagaagatgctgatcgagaagctgaattacctggtgttt3120
aaggataacgagttcgacaagaccggaggcgtgctgagggcataccagctgaccgccccc3180
tttgagacattcaagaagatgggcaagcagacaggcatcatctactatgtgccagccggc3240
ttcacctccaagatctgccccgtgacaggctttgtgaaccagctgtaccctaagtatgag3300
tccgtgtctaagagccaggagtttttcagcaagttcgataagatctgttataatctggac3360
aagggctacttcgagttttccttcgattataagaactttggcgacaaggccgccaagggc3420
aagtggaccatcgcctctttcggcagccggctgatcaactttagaaattccgataagaac3480
cacaattgggacacccgggaggtgtacccaacaaaggagctggagaagctgctgaaggac3540
tacagcatcgagtatggccacggcgagtgcatcaaggccgccatctgtggcgagagcgat3600
aagaagtttttcgccaagctgacctccgtgctgaatacaatcctgcagatgcggaacagc3660
aagaccggcacagagctggactacctgatctcccccgtggccgatgtgaacggcaacttc3720
ttcgacagcagacaggcccccaagaatatgcctcaggatgccgacgccaacggcgcctat3780
cacatcggcctgaagggcctgatgctgctgggcaggatcaagaacaatcaggagggcaag3840
aagctgaacctggtcatcaagaacgaggagtactttgagttcgtgcagaaccgcaacaat3900
tga3903
<210>5
<211>3903
<212>dna
<213>artificialsequence
<220>
<223>ep16的核苷酸序列
<400>5
atgagcatctatcaggagttcgtgaataagtacagcctgtccaagaccctgcggtttgag60
ctgatcccccagggcaagacactggagaacatcaaggccaggggcctgatcctggacgat120
gagaagcgcgccaaggactataagaaggccaagcagatcatcgataagtaccaccagttc180
tttatcgaggagatcctgagcagcgtgtgcatctctgaggatctgctgcagaattacagc240
gacgtgtatttcaagctgaagaagtctgacgatgacaacctgcagaaggacttcaagagc300
gccaaggacaccatcaagaagcagatcagcgagtatatcaaggactccgagaagtttaag360
aatctgttcaaccagaatctgatcgatgccaagaagggccaggagtccgacctgatcctg420
tggctgaagcagtctaaggacaatggcatcgagctgttcaaggccaactctgatatcacc480
gatatcgacgaggccctggagatcatcaagagctttaagggctggaccacatactttagc540
ggcttccacgagaacaggaagaacgtgtacagcagcaacgacatccctacaagcatcatc600
taccgcatcgtggatgacaatctgccaaagttcctggagaacaaggccaagtatgagtcc660
ctgaaggacaaggcccccgaggccatcaattacgagcagatcaagaaggatctggccgag720
gagctgaccttcgatatcgactataagacatccgaggtgaaccagcgggtgttttctctg780
gacgaggtgtttgagatcgccaatttcaacaattacctgaaccagtccggcatcaccaag840
ttcaatacaatcatcggcggcaagtttgtgaacggcgagaataccaagagaaagggcatc900
aacgagtacatcaatctgtatagccagcagatcaacgacaagaccctgaagaagtacaag960
atgagcgtgctgttcaagcagatcctgtccgatacagagtctaagagctttgtgatcgat1020
aagctggaggatgactctgacgtggtgaccacaatgcagagcttttatgagcagatcgcc1080
gccttcaagaccgtggaggagaagtctatcaaggagacactgagcctgctgttcgatgac1140
ctgaaggcccagaagctggacctgtctaagatctacttcaagaacgataagtccctgacc1200
gacctgtctcagcaggtgtttgatgactatagcgtgatcggcaccgccgtgctggagtac1260
atcacacagcagatcgccccaaagaacctggataatccctctaagaaggagcaggagctg1320
atcgccaagaagaccgagaaggccaagtatctgagcctggagacaatcaagctggccctg1380
gaggagttcaataagcaccgggatatcgacaagcagtgcagatttgaggagatcctggcc1440
aacttcgccgccatccccatgatctttgatgagatcgcccagaacaaggacaatctggcc1500
cagatctccatcaagtaccagaaccagggcaagaaggacctgctgcaggcctctgccgag1560
gatgacgtgaaggccatcaaggatctgctggaccagaccaacaatctgctgcacaagctg1620
aagatcttccacatctcccagtctgaggataaggccaatatcctggataaggacgagcac1680
ttttatctggtgttcgaggagtgttacttcgagctggccaacatcgtgcccctgtacaac1740
aagatcagaaattatatcacacagaagccttactccgacgagaagtttaagctgaacttc1800
gagaacagcaccctggccagaggctgggataagaatgtggagcctaacagaacagccatc1860
ctgttcatcaaggatgacaagtactatctgggcgtgatgaataagaagaacaataagatc1920
ttcgatgacaaggccatcaaggagaacaagggcgagggctacaagaagatcgtgtatagg1980
ctgctgcccggcgccaataagatgctgcctaaggtgttcttttccgccaagtctatcaag2040
ttctacaacccatccgaggacatcctgcggatcagaaatcactccacccacacaaagaac2100
ggctctccccagaagggctatgagaagtttgagttcaatatcgaggattgccggaagttt2160
atcgacttctacaagcagagcatctccaagcaccctgagtggaaggattttggcttcagg2220
tttagcgacacccagcggtacaactccatcgacgagttctacagagaggtggagaatcag2280
ggctataagctgacatttgagaacatctctgagagctacatcgacagcgtggtgaatcag2340
ggcaagctgtacctgttccagatctataacaaggacttcagcgcctattccaagggccgg2400
ccaaacctgcacaccctgtactggaaggccctgttcgatgagagaaatctgcaggacgtg2460
gtgtataagctgaacggcgaggccgagctgttttacaggaagcagtccatccctaagaag2520
atcacacacccagccaaggaggccatcgccaacaagaataaggacaatcctaagaaggag2580
agcgtgttcgagtacgatctgatcaaggacaagcggttcaccgaggataagttctttttc2640
cactgtccaatcacaatcaacttcaagtcctctggcgccaacaagtttaatgacgagatc2700
aatctgctgctgaaggagaaggccaacgatgtgcacatcctgagcatcgaccggggcgag2760
agacacctggcctactataccctggtggatggcaagggcaatatcatcaagcaggatacc2820
ttcaacatcatcggcaatgacaggatgaagacaaactaccacgataagctggccgccatc2880
gagaaggatagggactccgcccgcaaggactggaagaagatcaacaatatcaaggagatg2940
aaggagggctatctgtctcaggtggtgcacgagatcgccaagctggtcatcgagtacaat3000
gccatcgtggtgttcgaggatctgaacttcggctttaagaggggccgctttaaggtggag3060
aagcaggtgtatcagaagctggagaagatgctgatcgagaagctgaattacctggtgttt3120
aaggataacgagttcgacaagaccggaggcgtgctgagggcataccagctgaccgccccc3180
tttgagacattcaagaagatgggcaagcagacaggcatcatctactatgtgccagccggc3240
ttcacctccaagatctgccccgtgacaggctttgtgaaccagctgtaccctaagtatgag3300
tccgtgtctaagagccaggagtttttcagcaagttcgataagatctgttataatctggac3360
aagggctacttcgagttttccttcgattataagaactttggcgacaaggccgccaagggc3420
aagtggaccatcgcctctttcggcagccggctgatcaactttagaaattccgataagaac3480
cacaattgggacacccgggaggtgtacccaacaaaggagctggagaagctgctgaaggac3540
tacagcatcgagtatggccacggcgagtgcatcaaggccgccatctgtggcgagagcgat3600
aagaagtttttcgccaagctgacctccgtgctgaatacaatcctgcagatgcggaacagc3660
aagaccggcacagagctggactacctgatctcccccgtggccgatgtgaacggcaacttc3720
ttcgacagcagacaggcccccaagaatatgcctcaggatgccgacgccaacggcgcctat3780
cacatcggcctgaagggcctgatgctgctgggcaggatcaagaacaatcaggagggcaag3840
aagctgaacctggtcatcaagaacgaggagtactttgagttcgtgcagaaccgcaacaat3900
tga3903
- 还没有人留言评论。精彩留言会获得点赞!