基于C2c1核酸酶的基因组编辑系统和方法
本申请为申请日为2018年11月2日、申请号为201811300251.6、发明名称为“基于c2c1核酸酶的基因组编辑系统和方法”的发明专利申请的分案申请。
本发明涉及基因工程领域。具体而言,本发明涉及基于c2c1核酸酶的基因组编辑系统和方法。本发明还涉及可与不同c2c1核酸酶组合用于基因组编辑的人工向导rna。
发明背景
随着crispr-cas(clusteredregularlyinterspacedshortpalindromicrepeats-crispr-associated蛋白)系统的出现,精确的基因组编辑由于其在基因治疗中的光明前景已经成为最令人关注的领域。到目前为止,已成功利用三种类型的crispr-cas系统以促进哺乳动物基因组工程,包括ii型cas9(cong,l.etal.science339,819-823(2013);mali,p.etal.science339,823-826(2013))、v-a型cpf1(zetsche,b.etal.cell163,759-771(2015))和v-b型c2c1。对于ii型和v型crispr-cas系统,向导rna和cas效应蛋白是靶dna识别和切割的两种核心成分(wright,a.v.,nunez,j.k.&doudna,j.a.cell164,29-44(2016);shmakov,s.etal.natrevmicrobiol15,169-182(2017))。以前的研究表明在密切相关的cas9系统(fonfara,i.etal.nucleicacidsres42,2577-2590(2014))以及cpf1系统(zetsche,b.etal.cell163,759-771(2015))中,双rna(crrna和tracrrna)和蛋白质组分是可互换的,并能初步优化(nishimasu,h.etal.cell156,935-949(2014);zalatan,j.g.etal.cell160,339-350(2015))。虽然许多新兴的crispr-cas系统和研究促进crispr-cas系统的广泛应用(wright,a.v.,nunez,j.k.&doudna,j.a.cell164,29-44(2016);shmakov,s.etal.natrevmicrobiol15,169-182(2017)),但对于如何重新设计甚至从头合成促酶基因组编辑系统仍然知之甚少。
v-b型crispr-c2c1系统是一种新兴的具有前景的基因工程技术。然而,可用于哺乳动物基因组编辑的c2c1却很少,大大限制了其应用。本领域仍然需要新的可用于哺乳动物基因组编辑的基于c2c1核酸酶的基因组编辑系统。
发明简述
在一方面,本发明提供了一种用于对细胞基因组中的靶序列进行定点修饰的基因组编辑系统,其包含以下i)至v)中至少一项:
i)c2c1蛋白或其变体,和向导rna;
ii)包含编码c2c1蛋白或其变体的核苷酸序列的表达构建体,和向导rna;
iii)c2c1蛋白或其变体,和包含编码向导rna的核苷酸序列的表达构建体;
iv)包含编码c2c1蛋白或其变体的核苷酸序列的表达构建体,和包含编码向导rna的核苷酸序列的表达构建体;
v)包含编码c2c1蛋白或其变体的核苷酸序列和编码向导rna的核苷酸序列的表达构建体;
其中所述向导rna能够与所述c2c1蛋白或其变体形成复合物,将所述c2c1蛋白直系同源物或其变体靶向所述细胞基因组中的靶序列。
在一些实施方案中,所述c2c1蛋白是来自alicyclobacillusacidiphilus的aac2c1蛋白、来自alicyclobacilluskakegawensis的akc2c1蛋白、来自alicyclobacillusmacrosporangiidus的amc2c1蛋白、来自bacillushisashii的bhc2c1蛋白、来自bacillus属的bsc2c1蛋白、来自bacillus属的bs3c2c1蛋白、来自desulfovibrioinopinatus的dic2c1蛋白、来自laceyellasediminis的lsc2c1蛋白、来自spirochaetesbacterium的sbc2c1蛋白、来自tuberibacilluscalidus的tcc2c1蛋白。例如,所述c2c1蛋白是来自alicyclobacillusacidiphilusnbrc100859的aac2c1蛋白、来自alicyclobacilluskakegawensisnbrc103104的akc2c1蛋白、来自alicyclobacillusmacrosporangiidusstraindsm17980的amc2c1蛋白、来自bacillushisashiistrainc4的bhc2c1蛋白、来自bacillus属nsp2.1的bsc2c1蛋白、来自bacillus属v3-13contig_40的bs3c2c1蛋白、来自desulfovibrioinopinatusdsm10711的dic2c1蛋白、来自laceyellasediminisstrainrha1的lsc2c1蛋白、来自spirochaetesbacteriumgwb1_27_13的sbc2c1蛋白、来自tuberibacilluscalidusdsm17572的tcc2c1蛋白。
在第二方面,本发明提供了一种对细胞基因组中的靶序列进行定点修饰的方法,包括将本发明的基因组编辑系统导入所述细胞。
在第三方面,本发明提供了一种治疗有需要的对象中的疾病的方法,包括向所述对象递送有效量的本发明的基因组编辑系统以修饰所述对象中与所述疾病相关的基因。
在第四方面,本发明提供了本发明的基因组编辑系统在制备用于治疗有需要的对象中的疾病的药物组合物中的用途,其中所述基因组编辑系统用于修饰所述对象中与所述疾病相关的基因。
在第五方面,本发明提供了用于本发明的方法的试剂盒,该试剂盒包括本发明的基因组编辑系统,以及使用说明。
在第六方面,本发明提供了一种用于治疗有需要的对象中的疾病的药物组合物,其包含本发明的基因组编辑系统和药学可接受的载体,其中所述基因组编辑系统用于修饰所述对象中与所述疾病相关的基因。
附图简述
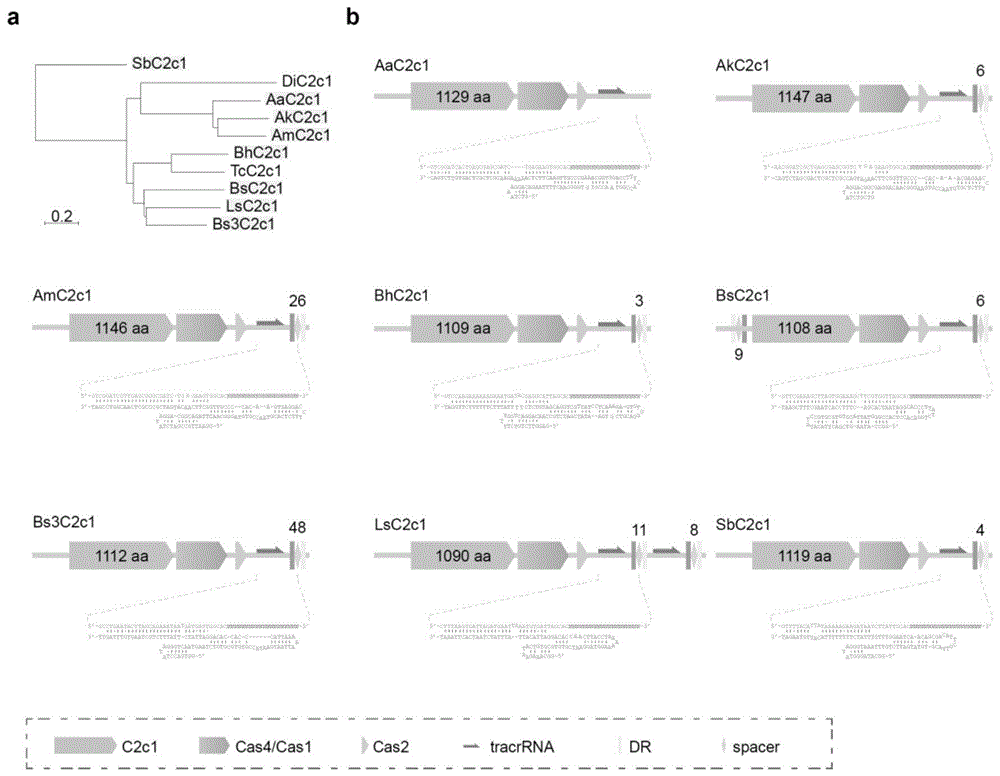
图1.选择用于基因组编辑测试的非冗余c2c1直系同源物的系统发生树及其基因座。
(a)邻接系统发生树,显示测试的c2c1直系同源物的进化关系。(b)对应于(a)中突出显示的8种c2c1蛋白的细菌基因座图谱crrnadr和推定的tracrrna的模拟共折叠显示出稳定的二级结构。dr,直接重复。每个细菌基因组间隔区(spacer)的数目在其crispr阵列的上方或下方表示。
图2.c2c1直系同源物的蛋白质比对:测试的10种c2c1直系同源物的氨基酸序列的多序列比对。保守的残基用红色背景突出显示,保守突变用轮廓和红色字体突出显示。
图3.人293t细胞中c2c1直系同源物介导的基因组靶向。
(a)t7ei测定结果表明在人类基因组中与其同源sgrna结合的八种c2c1蛋白的基因组靶向活性。三角形表示切割的条带。(b)t7ei测定结果表明在人293t细胞中由与其同源sgrna(bs3sgrna)结合的bs3c2c1介导的同时多重基因组靶向。(c)sanger测序显示由与bs3sgrna结合的bs3c2c1诱导的代表性插入缺失(indel)。pam和原间隔区序列分别用红色和蓝色着色。插入缺失和插入分别用紫色破折号和绿色小写字符表示。
图4.用于rna指导的基因组编辑的c2c1蛋白。
(a)本发明中测试的10种c2c1直系同源物的图形概述。示出其大小(氨基酸数目)。(b)t7ei测定结果表明在人293t细胞中由其同源sgrna指导的八种c2c1直系同源物的基因组靶向活性。三角形表示切割的条带。(c-d)t7ei测定结果表明在人293t细胞中由aasgrna(c)和aksgrna(d)指导的八种c2c1直系同源物的基因组靶向活性。三角形表示切割的条带。
图5.c2c1的sgrna的dna比对:测试衍生自10个c2c1基因座的8种sgrna的dna序列的多序列比对。
图6.不同c2c1直系同源物与sgrna之间的可互换性。
t7ei测定结果表明在人293t细胞中由aasgrna(a)、aksgrna(b)、amsgrna(c)、bs3sgrna(d)和lssgrna(e)指导的八种c2c1直系同源物的基因组靶向活性。红色三角形表示切割的条带。
图7.人工sgrna介导的多重基因组靶向。
(a)对应于dic2c1和tcc2c1的细菌基因座的图谱。两个c2c1基因座没有crispr阵列。(b-c)t7ei测定结果表明在人293t细胞中由aasgrna(b)和aksgrna(c)指导的aac2c1、dic2c1和tcc2c1的基因组靶向活性。三角形表示切割的条带。(d)t7ei测定结果表明在人293t细胞中由与aksgrna结合的tcc2c1介导的同时多重基因组靶向。(e)示意图说明人工sgrna支架13(artgrna13)的二级结构。(f)t7ei测定结果表明在人293t细胞中由与artgrna13结合的tcc2c1介导的同时多重基因组靶向。
图8.不同sgrna指导c2c1进行基因组编辑。
t7ei测定结果表明在人293t细胞中由aasgrna(a)、aksgrna(b)、amsgrna(c)、bs3sgrna(d)和lssgrna(e)指导的aac2c1、dic2c1和tcc2c1的基因组靶向活性。三角形表示切割的条带。
图9.tcc2c1介导的多重基因组编辑。
(a)t7ei测定结果表明在人293t细胞中由与amsgrna结合的tcc2c1介导的同时多重基因组靶向。(b-c)sanger测序显示由与aksgrna(b)和amsgrna(c)结合的tcc2c1诱导的代表性插入缺失。pam和原间隔区序列分别用红色和蓝色着色。插入缺失和插入分别用紫色破折号和绿色小写字符表示。
图10.人工sgrna指导tcc2c1进行基因组编辑。
(a)示意图说明36种人工sgrna(artgrna)支架(支架:1-12和14-37)的二级结构。(b)t7ei测定结果表明在人293t细胞中artsgrna指导的tcc2c1的基因组靶向活性。三角形表示切割的条带。(c)t7ei测定结果表明在人293t细胞中由与artgrna13结合的aac2c1介导的同时多重基因组靶向。
发明详述
一、定义
在本发明中,除非另有说明,否则本文中使用的科学和技术名词具有本领域技术人员所通常理解的含义。并且,本文中所用的蛋白质和核酸化学、分子生物学、细胞和组织培养、微生物学、免疫学相关术语和实验室操作步骤均为相应领域内广泛使用的术语和常规步骤。例如,本发明中使用的标准重组dna和分子克隆技术为本领域技术人员熟知,并且在如下文献中有更全面的描述:sambrook,j.,fritsch,e.f.和maniatis,t.,molecularcloning:alaboratorymanual;coldspringharborlaboratorypress:coldspringharbor,1989(下文称为“sambrook”)。同时,为了更好地理解本发明,下面提供相关术语的定义和解释。
在一方面,本发明提供了一种用于对细胞基因组中的靶序列进行定点修饰的基因组编辑系统,其包含以下i)至v)中至少一项:
i)c2c1蛋白或其变体,和向导rna;
ii)包含编码c2c1蛋白或其变体的核苷酸序列的表达构建体,和向导rna;
iii)c2c1蛋白或其变体,和包含编码向导rna的核苷酸序列的表达构建体;
iv)包含编码c2c1蛋白或其变体的核苷酸序列的表达构建体,和包含编码向导rna的核苷酸序列的表达构建体;
v)包含编码c2c1蛋白或其变体的核苷酸序列和编码向导rna的核苷酸序列的表达构建体;
其中所述向导rna能够与所述c2c1蛋白或其变体形成复合物,将所述c2c1蛋白或其变体靶向所述细胞基因组中的靶序列。在一些实施方案中,所述靶向导致所述靶序列中的一或多个核苷酸的取代、缺失和/或添加。
“基因组”如本文所用不仅涵盖存在于细胞核中的染色体dna,而且还包括存在于细胞的亚细胞组分(如线粒体、质体)中的细胞器dna。
“c2c1核酸酶”、“c2c1蛋白”和“c2c1”在本文中可互换使用,指的是包括c2c1蛋白或其片段的rna指导的核酸酶。c2c1具有向导rna介导的dna结合活性以及dna切割活性,能在向导rna的指导下靶向并切割dna靶序列形成dna双链断裂(dsb)。dsb能够激活细胞内固有的修复机制非同源末端连接(non-homologousendjoining,nhej)和同源重组(homologousrecombination,hr)对细胞中的dna损伤进行修复,在修复过程中,对该特定的dna序列进行定点编辑。
在一些实施方案中,所述c2c1蛋白是来自alicyclobacillusacidiphilus的aac2c1蛋白、来自alicyclobacilluskakegawensis的akc2c1蛋白、来自alicyclobacillusmacrosporangiidus的amc2c1蛋白、来自bacillushisashii的bhc2c1蛋白、来自bacillus属的bsc2c1蛋白、来自bacillus属的bs3c2c1蛋白、来自desulfovibrioinopinatus的dic2c1蛋白、来自laceyellasediminis的lsc2c1蛋白、来自spirochaetesbacterium的sbc2c1蛋白、来自tuberibacilluscalidus的tcc2c1蛋白。
例如,所述c2c1蛋白是来自alicyclobacillusacidiphilusnbrc100859的aac2c1蛋白、来自alicyclobacilluskakegawensisnbrc103104的akc2c1蛋白、来自alicyclobacillusmacrosporangiidusstraindsm17980的amc2c1蛋白、来自bacillushisashiistrainc4的bhc2c1蛋白、来自bacillus属nsp2.1的bsc2c1蛋白、来自bacillus属v3-13contig_40的bs3c2c1蛋白、来自desulfovibrioinopinatusdsm10711的dic2c1蛋白、来自laceyellasediminisstrainrha1的lsc2c1蛋白、来自spirochaetesbacteriumgwb1_27_13的sbc2c1蛋白、来自tuberibacilluscalidusdsm17572的tcc2c1蛋白。
在本发明一些实施方式中,所述c2c1蛋白是其天然基因座不具有crispr阵列的c2c1蛋白。在一些实施方式中,所述天然基因座不具有crispr阵列的c2c1蛋白是dic2c1或tcc2c1蛋白。
在一些实施方案中,所述c2c1蛋白包含seqidno:1-10中任一所示的氨基酸序列。例如,所述aac2c1、akc2c1、amc2c1、bhc2c1、bsc2c1、bs3c2c1、dic2c1、lsc2c1、sbc2c1、tcc2c1蛋白分别包含seqidno:1-10所示氨基酸序列。
在一些实施方案中,所述c2c1蛋白的变体分别包含与野生型c2c1蛋白(如野生型aac2c1、akc2c1、amc2c1、bhc2c1、bsc2c1、bs3c2c1、dic2c1、lsc2c1、sbc2c1或tcc2c1蛋白)具有至少80%、至少85%、至少90%、至少95%、至少96%、至少97%、至少98%、至少99%序列相同性的氨基酸序列,并且分别具有野生型c2c1蛋白(如野生型aac2c1、akc2c1、amc2c1、bhc2c1、bsc2c1、bs3c2c1、dic2c1、lsc2c1、sbc2c1、tcc2c1蛋白)的基因组编辑和/靶向活性。
在一些实施方案中,所述c2c1蛋白的变体分别包含相对于野生型c2c1蛋白(如野生型aac2c1、akc2c1、amc2c1、bhc2c1、bsc2c1、bs3c2c1、dic2c1、lsc2c1、sbc2c1、tcc2c1蛋白)具有一或多个氨基酸残基取代、缺失或添加的氨基酸序列,并且分别具有野生型c2c1蛋白(如野生型aac2c1、akc2c1、amc2c1、bhc2c1、bsc2c1、bs3c2c1、dic2c1、lsc2c1、sbc2c1、tcc2c1蛋白)的基因组编辑和/或靶向活性。例如,所述c2c1蛋白的变体分别包含相对于野生型c2c1蛋白(如野生型aac2c1、akc2c1、amc2c1、bhc2c1、bsc2c1、bs3c2c1、dic2c1、lsc2c1、sbc2c1、tcc2c1蛋白)具有1个、2个、3个、4个、5个、6个、7个、8个、9个或10个氨基酸残基取代、缺失或添加的氨基酸序列。在一些实施方案中,所述氨基酸取代是保守型取代。
“多肽”、“肽”、和“蛋白质”在本发明中可互换使用,指氨基酸残基的聚合物。该术语适用于其中一个或多个氨基酸残基是相应的天然存在的氨基酸的人工化学类似物的氨基酸聚合物,以及适用于天然存在的氨基酸聚合物。术语“多肽”、“肽”、“氨基酸序列”和“蛋白质”还可包括修饰形式,包括但不限于糖基化、脂质连接、硫酸盐化、谷氨酸残基的γ羧化、羟化和adp-核糖基化。
序列“相同性”具有本领域公认的含义,并且可以利用公开的技术计算两个核酸或多肽分子或区域之间序列相同性的百分比。可以沿着多核苷酸或多肽的全长或者沿着该分子的区域测量序列相同性。(参见,例如:computationalmolecularbiology,lesk,a.m.,ed.,oxforduniversitypress,newyork,1988;biocomputing:informaticsandgenomeprojects,smith,d.w.,ed.,academicpress,newyork,1993;computeranalysisofsequencedata,parti,griffin,a.m.,andgriffin,h.g.,eds.,humanapress,newjersey,1994;sequenceanalysisinmolecularbiology,vonheinje,g.,academicpress,1987;andsequenceanalysisprimer,gribskov,m.anddevereux,j.,eds.,mstocktonpress,newyork,1991)。虽然存在许多测量两个多核苷酸或多肽之间的相同性的方法,但是术语“相同性”是技术人员公知的(carrillo,h.&lipman,d.,siamjappliedmath48:1073(1988))。
在肽或蛋白中,合适的保守型氨基酸取代是本领域技术人员已知的,并且一般可以进行而不改变所得分子的生物活性。通常,本领域技术人员认识到多肽的非必需区中的单个氨基酸取代基本上不改变生物活性(参见,例如,watsonetal.,molecularbiologyofthegene,4thedition,1987,thebenjamin/cummingspub.co.,p.224)。
在一些实施方案中,所述c2c1蛋白的变体包含核酸酶死亡的c2c1蛋白(dc2c1)。核酸酶死亡的c2c1蛋白指的是保留向导rna介导的dna结合活性但是不具备双链dna切割活性的c2c1蛋白。在一些实施方案中,所述核酸酶死亡的c2c1蛋白涵盖c2c1切口酶,其只切割双链靶dna的一条链。
在一些实施方案中,所述c2c1蛋白的变体是核酸酶死亡的c2c1蛋白与脱氨酶的融合蛋白。例如,所述融合蛋白中的核酸酶死亡的c2c1蛋白与脱氨酶可以通过接头例如肽接头连接。
如本发明所用,“脱氨酶”是指催化脱氨基反应的酶。在本发明一些实施方式中,所述脱氨酶指的是胞嘧啶脱氨酶,其能够接受单链dna作为底物并能够催化胞苷或脱氧胞苷分别脱氨化为尿嘧啶或脱氧尿嘧啶。在本发明一些实施方式中,所述脱氨酶指的是腺嘌呤脱氨酶,其能够接受单链dna作为底物并能够催化腺苷或脱氧腺苷(a)形成肌苷(i)。通过使用核酸酶死亡的c2c1蛋白与脱氨酶的融合蛋白,可以实现靶dna序列中的碱基编辑,例如c至t的转换或a至g的转换。
在本发明的一些实施方案中,本发明的基因组编辑系统中的c2c1蛋白或其变体还可以包含核定位序列(nls)。一般而言,所述c2c1蛋白或其变体中的一个或多个nls应具有足够的强度,以便在细胞核中驱动所述c2c1蛋白或其变体以可实现其基因编辑功能的量积聚。一般而言,核定位活性的强度由所述c2c1蛋白或其变体中nls的数目、位置、所使用的一个或多个特定的nls、或这些因素的组合决定。
在本发明的一些实施方案中,本发明的基因组编辑系统中的c2c1蛋白或其变体的nls可以位于n端和/或c端。在一些实施方案中,所述c2c1蛋白或其变体包含约1、2、3、4、5、6、7、8、9、10个或更多个nls。在一些实施方案中,所述c2c1蛋白或其变体包含在或接近于n端的约1、2、3、4、5、6、7、8、9、10个或更多个nls。在一些实施方案中,所述c2c1蛋白或其变体包含在或接近于c端约1、2、3、4、5、6、7、8、9、10个或更多个nls。在一些实施方案中,所述c2c1蛋白或其变体包含这些的组合,如包含在n端的一个或多个nls以及在c端的一个或多个nls。当存在多于一个nls时,每一个可以被选择为不依赖于其他nls。在本发明的一些实施方式中,所述c2c1蛋白或其变体包含2个nls,例如所述2个nls分别位于n端和c端。
一般而言,nls由暴露于蛋白表面上的带正电的赖氨酸或精氨酸的一个或多个短序列组成,但其他类型的nls也是已知的。nls的非限制性实例包括:kkrkv、pkkkrkv,或sggspkkkrkv。
此外,根据所需要编辑的dna位置,本发明的c2c1蛋白或其变体还可以包括其他的定位序列,例如细胞质定位序列、叶绿体定位序列、线粒体定位序列等。
在本发明的一些实施方案中,所述靶序列长度为18-35个核苷酸,优选20个核苷酸。在本发明的一些实施方案中,所述靶序列在其5’端侧翼为选自:5’tttn-3’、5’attn-3’、5’gttn-3’、5’cttn-3’、5’ttc-3’、5’ttg-3’、5’tta-3’、5’ttt-3’、5’tan-3’、5’tgn-3’、5’tcn-3’和5’atc-3’的pam(前间区邻近基序)序列,其中n选自a、g、c和t。
在本发明中,待进行修饰的靶序列可以位于基因组的任何位置,例如位于功能基因如蛋白编码基因内,或者例如可以位于基因表达调控区如启动子区或增强子区,从而实现对所述基因功能修饰或对基因表达的修饰。可以通过t7ei、pcr/re或测序方法检测基因组靶序列中的取代、缺失和/或添加。
“向导rna”和“grna”在本文中可互换使用,通常由部分互补形成复合物的crrna和tracrrna分子构成,其中crrna包含与靶序列具有足够相同性以便与靶序列的互补序列杂交并且指导crispr复合物(c2c1+crrna+tracrrna)与该靶序列以序列特异性方式结合的序列。然而,可以设计并使用单向导rna(sgrna),其同时包含crrna和tracrrna的特征。
在本发明的一些实施方案中,所述向导rna是sgrna。在一些具体实施方案中,所述sgrna由选自以下之一的核酸序列编码:
5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagcgagcgatctgagaagtggcac-nx-3’(aasgrna);
5’-tcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaagatgaccgctcgctcagcgatctgacaacggatcgctgagcgagcggtctgagaagtggcac-nx-3’(aksgrna1);
5’-ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttggcttcaacatgatcgcccgctcaacggtccgatgtcggatcgttgagcgggcgatctgagaagtggcac-nx-3’(amsgrna1);
5’-gaggttctgtcttttggtcaggacaaccgtctagctataagtgctgcagggtgtgagaaactcctattgctggacgatgtctcttttatttcttttttcttggatgtccaagaaaaaagaaatgatacgaggcattagcac-nx-3’(bhsgrna);
5’-ccataagtcgacttacatatccgtgcgtgtgcattatgggcccatccacaggtctattcccacggataatcacgactttccactaagctttcgaatgttcgaaagcttagtggaaagcttcgtggttagcac-nx-3’(bssgrna);
5’-ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggattatcttatttctgctaagtgtttagttgcctgaatacttagcagaaataatgatgattggcac-nx-3’(bs3sgrna);
5’-ggcaaagaatactgtgcgtgtgctaaggatggaaaaaatccattcaaccacaggattacattatttatctaatcacttaaatctttaagtgattagatgaattaaatgtgattagcac-nx-3’(lssgrna);或
5’-gtcttagggtatatcccaaatttgtcttagtatgtgcattgcttacagcgacaactaaggtttgtttatcttttttttacattgtaagatgttttacattataaaaagaagataatcttattgcac-nx-3’(sbsgrna);
其中nx表示x个连续的核苷酸组成的核苷酸序列(spacer序列),n各自独立地选自a、g、c和t;x为18≤x≤35的整数。优选地,x=20。在一些实施方案中,序列nx(spacer序列)能够与靶序列的互补序列特异性杂交。所述sgrna中除nx之外的序列为sgrna的支架(scaffold)序列。在一些实施方案中,所述sgrna包含由seqidno:31-38中任一项的核苷酸序列编码的支架序列。
本发明令人惊奇地发现,不同的c2c1系统中的c2c1蛋白以及向导rna可以互换使用,从而使得可以人工设计通用的向导rna。
因此在另一方面,本发明提供一种人工sgrna,其由选自以下的核苷酸序列编码:
5’-ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’(artsgrna1);
5’-ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna2);
5’-ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacagacttcaagcgaagtggcac-nx-3’(artsgrna3);
5’-ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’(artsgrna4);
5’-ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna5);
5’-ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccaccacagacttcaagcgaagtggcac-nx-3’(artsgrna6);
5’-ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’(artsgrna7);
5’-ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna8);
5’-ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccaccacagacttcaagcgaagtggcac-nx-3’(artsgrna9);
5’-ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagcttcaaagaagtggcac-nx-3’(artsgrna10);
5’-ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna11);
5’-ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggcttcaaagaagtggcac-nx-3’(artsgrna12);
5’-ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagcttcaaagaagtggcac-nx-3’(artsgrna13);
5’-ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna14);
5’-ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggcttcaaagaagtggcac-nx-3’(artsgrna15);
5’-ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagcttcaaagaagtggcac-nx-3’(artsgrna16);
5’-ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna17);
5’-ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggcttcaaagaagtggcac-nx-3’(artsgrna18);
5’-ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagattatctatgatgattggcac-nx-3’(artsgrna19);
5’-ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggattatctatgatgattggcac-nx-3’(artsgrna20);
5’-ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggattatctatgatgattggcac-nx-3’(artsgrna21);
5’-ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagattatctatgatgattggcac-nx-3’(artsgrna22);
5’-ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggattatctatgatgattggcac-nx-3’(artsgrna23);
5’-ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggattatctatgatgattggcac-nx-3’(artsgrna24);
5’-ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagattatctatgatgattggcac-nx-3’(artsgrna25);
5’-ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggattatctatgatgattggcac-nx-3’(artsgrna26);
5’-ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggattatctatgatgattggcac-nx-3’(artsgrna27);
5’-ggtctaaaggacagaacaacgggatgtgccaatgcactctttccaggagtgaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna28);
5’-ggtcgtctataggacggcgagcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna29);
5’-ggaattgccgatctataggacggcagatttttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’(artsgrna30);
5’-ggaattgccgatctataggacggcagattgacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna31);
5’-ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna32);
5’-ggtctaaaggacagaacaacgggatgtgccaatgcactctttccaggagtgaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna33);
5’-ggtcgtctataggacggcgagcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna34);
5’-ggaattgccgatctataggacggcagatttttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagcttcaaagaagtggcac-nx-3’(artsgrna35);
5’-ggaattgccgatctataggacggcagattgacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrn36a);或
5’-ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna37),
其中nx表示x个连续的核苷酸组成的核苷酸序列(spacer序列),n各自独立地选自a、g、c和t;x为18≤x≤35的整数。优选地,x=20。在一些实施方案中,序列nx(spacer序列)能够与靶序列的互补序列特异性杂交。所述sgrna中除nx之外的序列为sgrna的支架(scaffold)序列。
在一些实施方案中,所述人工sgrna包含由seqidno:39-75中任一项的核苷酸序列编码的支架序列。
在一些实施方案中,本发明的基因组编辑系统中的向导rna是本发明的人工sgrna。
为了在靶细胞中获得有效表达,在本发明的一些实施方式中,所述编码c2c1蛋白或其变体的核苷酸序列针对待进行基因组编辑的细胞所来自的生物体进行密码子优化。
密码子优化是指通过用在宿主细胞的基因中更频繁地或者最频繁地使用的密码子代替天然序列的至少一个密码子(例如约或多于约1、2、3、4、5、10、15、20、25、50个或更多个密码子同时维持该天然氨基酸序列而修饰核酸序列以便增强在感兴趣宿主细胞中的表达的方法。不同的物种对于特定氨基酸的某些密码子展示出特定的偏好。密码子偏好性(在生物之间的密码子使用的差异)经常与信使rna(mrna)的翻译效率相关,而该翻译效率则被认为依赖于被翻译的密码子的性质和特定的转运rna(trna)分子的可用性。细胞内选定的trna的优势一般反映了最频繁用于肽合成的密码子。因此,可以将基因定制为基于密码子优化在给定生物中的最佳基因表达。密码子利用率表可以容易地获得,例如在www.kazusa.orjp/codon/上可获得的密码子使用数据库(“codonusagedatabase”)中,并且这些表可以通过不同的方式调整适用。参见,nakamuray.等,“codonusagetabulatedfromtheinternationaldnasequencedatabases:statusfortheyear2000.nucl.acidsres.,28:292(2000)。
可通过本发明的系统进行基因组编辑的细胞所来自的生物体优选是真核生物,包括但不限于,哺乳动物如人、小鼠、大鼠、猴、犬、猪、羊、牛、猫;家禽如鸡、鸭、鹅;植物包括单子叶植物和双子叶植物,例如水稻、玉米、小麦、高粱、大麦、大豆、花生、拟南芥等。
在本发明的一些具体实施方式中,所述编码c2c1蛋白或其变体的核苷酸序列针对人进行密码子优化。在一些具体实施方式中,所述密码子优化的编码aac2c1、akc2c1、amc2c1、bhc2c1、bsc2c1、bs3c2c1、dic2c1、lsc2c1、sbc2c1、tcc2c1蛋白的核苷酸序列分别选自seqidno:11-20。
根据本发明的一些实施方式,本发明所述系统的表达构建体中所述编码c2c1蛋白或其变体的核苷酸序列和/或所述编码向导rna的核苷酸序列与表达调控元件如启动子可操作地连接。
如本发明所用,“表达构建体”是指适于感兴趣的核苷酸序列在生物体中表达的载体如重组载体。“表达”指功能产物的产生。例如,核苷酸序列的表达可指核苷酸序列的转录(如转录生成mrna或功能rna)和/或rna翻译成前体或成熟蛋白质。本发明的“表达构建体”可以是线性的核酸片段、环状质粒、病毒载体,或者,在一些实施方式中,可以是能够翻译的rna(如mrna)。
本发明的“表达构建体”可包含不同来源的调控序列和感兴趣的核苷酸序列,或相同来源但以不同于通常天然存在的方式排列的调控序列和感兴趣的核苷酸序列。
“调控序列”和“调控元件”可互换使用,指位于编码序列的上游(5'非编码序列)、中间或下游(3'非编码序列),并且影响相关编码序列的转录、rna加工或稳定性或者翻译的核苷酸序列。调控序列可包括但不限于启动子、翻译前导序列、内含子和多腺苷酸化识别序列。
“启动子”指能够控制另一核酸片段转录的核酸片段。在本发明的一些实施方案中,启动子是能够控制细胞中基因转录的启动子,无论其是否来源于所述细胞。启动子可以是组成型启动子或组织特异性启动子或发育调控启动子或诱导型启动子。
“组成型启动子”指一般将引起基因在多数细胞类型中在多数情况下表达的启动子。“组织特异性启动子”和“组织优选启动子”可互换使用,并且指主要但非必须专一地在一种组织或器官中表达,而且也可在一种特定细胞或细胞型中表达的启动子。“发育调控启动子”指其活性由发育事件决定的启动子。“诱导型启动子”响应内源性或外源性刺激(环境、激素、化学信号等)而选择性表达可操纵连接的dna序列。
如本文中所用,术语“可操作地连接”指调控元件(例如但不限于,启动子序列、转录终止序列等)与核酸序列(例如,编码序列或开放读码框)连接,使得核苷酸序列的转录被所述转录调控元件控制和调节。用于将调控元件区域可操作地连接于核酸分子的技术为本领域已知的。
本发明可使用的启动子的实例包括但不限于聚合酶(pol)i、polii或poliii启动子。poli启动子的实例包括鸡rnapoli启动子。polii启动子的实例包括但不限于巨细胞病毒立即早期(cmv)启动子、劳斯肉瘤病毒长末端重复(rsv-ltr)启动子和猿猴病毒40(sv40)立即早期启动子。poliii启动子的实例包括u6和h1启动子。可以使用诱导型启动子如金属硫蛋白启动子。启动子的其他实例包括t7噬菌体启动子、t3噬菌体启动子、β-半乳糖苷酶启动子和sp6噬菌体启动子。当用于植物时,启动子可以是花椰菜花叶病毒35s启动子、玉米ubi-1启动子、小麦u6启动子、水稻u3启动子、玉米u3启动子、水稻肌动蛋白启动子。
可通过本发明的系统进行基因组编辑的细胞优选是真核生物细胞,包括但不限于,哺乳动物细胞如人、小鼠、大鼠、猴、犬、猪、羊、牛、猫;家禽如鸡、鸭、鹅的细胞;植物细胞包括单子叶植物细胞和双子叶植物细胞,例如水稻、玉米、小麦、高粱、大麦、大豆、花生、拟南芥等的细胞。在本发明的一些实施方案中,所述细胞是真核生物细胞,优选哺乳动物细胞,更优选是人细胞。
在另一方面,本发明提供了一种修饰细胞基因组中靶序列的方法,包括将本发明的基因组编辑系统导入所述细胞,由此所述向导rna将所述c2c1蛋白或其变体靶向所述细胞基因组中的靶序列。在一些实施方案中,所述靶向导致所述靶序列中的一或多个核苷酸的取代、缺失和/或添加。
将本发明的基因组编辑系统的核酸分子(例如质粒、线性核酸片段、rna等)或蛋白质“导入”细胞是指用所述核酸或蛋白质转化细胞,使得所述核酸或蛋白质在细胞中能够发挥功能。本发明所用的“转化”包括稳定转化和瞬时转化。“稳定转化”指将外源核苷酸序列导入基因组中,导致外源基因稳定遗传。一旦稳定转化,外源核酸序列稳定地整合进所述生物体和其任何连续世代的基因组中。“瞬时转化”指将核酸分子或蛋白质导入细胞中,执行功能而没有外源基因稳定遗传。瞬时转化中,外源核酸序列不整合进基因组中。
可用于将本发明的基因组编辑系统导入细胞的方法包括但不限于:磷酸钙转染、原生质融合、电穿孔、脂质体转染、微注射、病毒感染(如杆状病毒、痘苗病毒、腺病毒、腺相关病毒、慢病毒和其他病毒)、基因枪法、peg介导的原生质体转化、土壤农杆菌介导的转化。
在一些实施方式中,所述方法在体外进行。例如,所述细胞是分离的细胞。在一些实施方式中,所述细胞是car-t细胞。在一些实施方式中,所述细胞是诱导的胚胎干细胞。
在另一些实施方式中,所述方法还可以在体内进行。例如,所述细胞是生物体内的细胞,可以通过例如病毒介导的方法将本发明的系统体内导入所述细胞。例如,所述细胞可以是患者体内的肿瘤细胞。
在另一方面,本发明还提一种产生经遗传修饰的细胞的方法,包括将本发明的基因组编辑系统导入细胞中,由此所述向导rna将所述c2c1蛋白或其变体靶向所述细胞基因组中的靶序列。在一些实施方式中,所述靶向导致所述靶序列中的一或多个核苷酸取代、缺失和/或添加。
在另一方面,本发明还提供经遗传修饰的生物体,其包含通过本发明的方法产生的经遗传修饰的细胞或其后代。
如本文所用,“生物体”包括适于基因组编辑的任何生物体,优选真核生物。生物体的实例包括但不限于,哺乳动物如人、小鼠、大鼠、猴、犬、猪、羊、牛、猫;家禽如鸡、鸭、鹅;植物包括单子叶植物和双子叶植物,例如水稻、玉米、小麦、高粱、大麦、大豆、花生、拟南芥等。在本发明的一些实施方案中,所述生物体是真核生物,优选哺乳动物,更优选人。
如本文所用,“经遗传修饰的生物体”或“经遗传修饰的细胞”意指在其基因组内包含外源多核苷酸或修饰的基因或表达调控序列的生物体或细胞。例如外源多核苷酸能够稳定地整合进生物体或细胞的基因组中,并遗传连续的世代。外源多核苷酸可单独地或作为重组dna构建体的部分整合进基因组中。修饰的基因或表达调控序列为在生物体或细胞基因组中所述序列包含单个或多个脱氧核苷酸取代、缺失和添加。针对序列而言的“外源”意指来自外来物种的序列,或者如果来自相同物种,则指通过蓄意的人为干预而从其天然形式发生了组成和/或基因座的显著改变的序列。
在另一方面,本发明提供了一种基因表达调控系统,其基于本发明的核酸酶死亡的c2c1蛋白。此系统尽管并没有改变靶基因的序列,在本文范围内也定义为基因组编辑系统。
在一些实施方案中,本发明的基因表达调控系统是基因抑制或沉默系统,其可以包含以下之一:
i)核酸酶死亡的c2c1蛋白或其与转录阻遏蛋白的融合蛋白,和向导rna;
ii)包含编码核酸酶死亡的c2c1蛋白或其与转录阻遏蛋白的融合蛋白的核苷酸序列的表达构建体,和向导rna;
iii)核酸酶死亡的c2c1蛋白或其与转录阻遏蛋白的融合蛋白,和包含编码向导rna的核苷酸序列的表达构建体;
iv)包含编码核酸酶死亡的c2c1蛋白或其与转录阻遏蛋白的融合蛋白的核苷酸序列的表达构建体,和包含编码向导rna的核苷酸序列的表达构建体;或
v)包含编码核酸酶死亡的c2c1蛋白或其与转录阻遏蛋白的融合蛋白的核苷酸序列和编码向导rna的核苷酸序列的表达构建体。
所述核酸酶死亡的c2c1蛋白或向导rna的定义如上所述。所述转录阻遏蛋白的选择属于本领域技术人员的技能范围。
如本文所用,基因抑制或沉默是指基因表达水平的下调或消除,优选在转录水平。
然而,本发明的基因表达调控系统还可以使用核酸酶死亡的c2c1蛋白和转录激活蛋白的融合蛋白。在此种情况下,所述基因表达调控系统是基因表达激活系统。例如,本发明的基因表达激活系统可以包含以下之一:
i)核酸酶死亡的c2c1蛋白和转录激活蛋白的融合蛋白,和向导rna;
ii)包含编码核酸酶死亡的c2c1蛋白和转录激活蛋白的融合蛋白的核苷酸序列的表达构建体,和向导rna;
iii)核酸酶死亡的c2c1蛋白和转录激活蛋白的融合蛋白,和包含编码向导rna的核苷酸序列的表达构建体;
iv)包含编码核酸酶死亡的c2c1蛋白和转录激活蛋白的融合蛋白的核苷酸序列的表达构建体,和包含编码向导rna的核苷酸序列的表达构建体;或
v)包含编码核酸酶死亡的c2c1蛋白和转录激活蛋白的融合蛋白的核苷酸序列和编码向导rna的核苷酸序列的表达构建体。
所述核酸酶死亡的c2c1蛋白或向导rna的定义如上所述。所述转录激活蛋白的选择属于本领域技术人员的技能范围。
如本文所用,基因激活是指基因表达水平的上调,优选在转录水平。
在另一方面,本发明还涵盖本发明的基因组编辑系统在疾病治疗中的应用。
通过本发明的基因组编辑系统对疾病相关基因进行修饰,可以实现疾病相关基因的上调、下调、失活、激活或者突变纠正等,从而实现疾病的预防和/或治疗。例如,本发明中靶序列可以位于疾病相关基因的蛋白编码区内,或者例如可以位于基因表达调控区如启动子区或增强子区,从而可以实现对所述疾病相关基因功能修饰或对疾病相关基因表达的修饰。
“疾病相关”基因是指与非疾病对照的组织或细胞相比,在来源于疾病影响的组织的细胞中以异常水平或以异常形式产生转录或翻译产物的任何基因。在改变的表达与疾病的出现和/或进展相关的情况下,它可以是以异常高的水平被表达的基因;它可以是以异常低的水平被表达的基因。疾病相关基因还指具有一个或多个突变或直接负责或与一个或多个负责疾病的病因学的基因连锁不平衡的遗传变异的基因。转录的或翻译的产物可以是已知的或未知的,并且可以处于正常或异常水平。
因此,本发明还提供治疗有需要的对象中的疾病的方法,包括向所述对象递送有效量的本发明的基因组编辑系统以修饰与所述疾病相关的基因。
本发明还提供本发明的基因组编辑系统在制备用于治疗有需要的对象中的疾病的药物组合物中的用途,其中所述基因组编辑系统用于修饰与所述疾病相关的基因。
本发明还提供用于治疗有需要的对象中的疾病的药物组合物,其包含本发明的基因组编辑系统和药学可接受的载体,其中所述基因组编辑系统用于修饰与所述疾病相关的基因。
在一些实施方式中,所述对象是哺乳动物,例如人。
所述疾病的实例包括但不限于肿瘤、炎症、帕金森病、心血管疾病、阿尔茨海默病、自闭症、药物成瘾、年龄相关性黄斑变性、精神分裂症、遗传性疾病等。
在仍另一方面,本发明的范围内还包括用于本发明的方法的试剂盒,该试剂盒包括本发明的基因组编辑系统,以及使用说明。试剂盒一般包括表明试剂盒内容物的预期用途和/或使用方法的标签。术语标签包括在试剂盒上或与试剂盒一起提供的或以其他方式随试剂盒提供的任何书面的或记录的材料。
实施例
为了便于理解本发明,下面将参照相关具体实施例及附图对本发明进行更全面的描述。附图中给出了本发明的较佳实施例。但是,本发明可以以许多不同的形式来实现,并不限于本文所描述的实施例。相反地,提供这些实施例的目的是使对本发明的公开内容的理解更加透彻全面。
材料与方法
1.dna操作
根据molecularcloning:alaboratorymanual并进行一些修改进行dna操作,包括dna制备、消化、连接、扩增、合成、纯化、琼脂糖凝胶电泳等。简而言之,通过连接退火的寡核苷酸(表1)到bsai消化的puc19-u6-sgrna(seqidno:23-30)载体中构建用于细胞转染测定的靶向sgrna。
表1.人类基因组靶靶序列。
2.从头基因合成和质粒构建
采用psi-blast程序(altschul,s.f.etal.nucleicacidsres25,3389-3402(1997))鉴定新的crispr-c2c1直系同源物。其编码序列进行人源化(grote,a.etal.,nucleicacidsres33,w526-531(2005)),并且使用genedesign程序(richardson,s.m.etal.,genomeres16,550-556(2006))设计用于c2c1基因和sgrna合成的寡核苷酸。根据文献(li,g.etal.,methodsmolbiol1073,9-17(2013))合成各c2c1基因。使用
3.细胞培养和转染
将人胚胎肾293t细胞在补充有10%胎牛血清(fbs,gibco)和1%抗生素-抗真菌剂(gibco)的dulbecco'smodifiedeagle培养基(dmem,gibco)中于37℃,5%co2孵育下培养。按照制造商推荐的方案,使用lipofectamineltx(invitrogen)转染293t细胞。对于48孔板的每个孔,使用总共400ng质粒(c2c1:sgrna=2:1)。然后在转染后48小时,直接收获细胞用于基因组dna提取。
4.t7核酸内切酶i(t7ei)测定和sanger测序
将收获的细胞直接用补充有蛋白酶k的bufferl(bimake)裂解,并在55℃下孵育3小时并在95℃下灭活10分钟。对每个基因的c2c1靶位点周围的基因组区域进行pcr扩增。将200-400ngpcr产物与ddh2o混合至终体积10μl,并根据先前方法进行再退火过程以形成异源双链体。重新退火后,用1/10体积的nebuffertm2.1和0.2μlt7ei(neb)在37℃处理产物30分钟,并在3%琼脂糖凝胶上进行分析。基于相对条带强度对插入缺失进行定量(cong,l.etal.,science339,819-823(2013))。将t7ei测定鉴定的突变产物克隆到ta克隆载体中,并转化到感受态大肠杆菌菌株(transgenbiotech)中。过夜培养后,随机挑出菌落并测序。
实施例1、新c2c1蛋白鉴定
选择并从头合成来自不同细菌的六种代表性c2c1蛋白,以及之前报道的四种c2c1直系同源物,在人胚胎肾293t细胞中进行基因组编辑(图1、2和seqidno:1-10)。在这10种c2c1直系同源物中,来自d.inopinatus(dic2c1)和t.calidus(tcc2c1)的c2c1既没有可预测的前体crisprrna(pre-crrna)也没有反式激活crrna(tracrrna)(图1b),提示这两种c2c1蛋白可能不适合基因组编辑应用。
为了进行哺乳动物基因组编辑,用单独的c2c1酶和其靶向含有适当pam的人内源基因座的同源单向导rna(sgrna)共转染293t细胞(图1)。t7核酸内切酶(t7ei)测定的结果显示,除了发明人先前已经鉴定的aac2c1和akc2c1之外,四种新的c2c1直系同源物(amc2c1、bhc2c1、bs3c2c1和lsc2c1)稳健地编辑人类基因组,尽管它们的靶向效率在不同的直系同源物之间和在不同的靶向位点不同(图1b和图3a)。还通过简单地使用多个sgrna,使用bs3c2c1实现多重基因组编辑,同时编辑人类基因组中的四个位点(图3b,c)。这些新发现的c2c1直系同源物扩展对基于c2c1的基因组编辑的选择。
实施例2、不同c2c1及双rna的可互换性
为了研究c2c1系统中双rna(crrna和tracrrna)和蛋白质组分之间的可互换性,首先分析c2c1蛋白和双rna两者的保守性。除了c2c1直系同源物的保守氨基酸序列外(图4a和图2),前体crrna:tracrrna双链体的dna序列及其二级结构也表现出高保守性(图1b和图5)。接下来,用分别与来自8个c2c1系统的各sgrna复合的8种c2c1直系同源物,在293t细胞中进行基因组编辑。如t7ei测定的结果所示,衍生自aac2c1、akc2c1、amc2c1、bs3c2c1和lsc2c1基因座的sgrna可以替代原始sgrna用于哺乳动物基因组编辑,尽管在不同c2c1直系同源物和sgrna之间的活性有所不同(图4c,d和图6)。这些结果证明不同c2c1和来自不同c2c1基因座的双rna之间的可互换性。
实施例3、利用天然基因座无crispr阵列的c2c1进行基因组编辑
本发明进一步选择两个基因座没有携带crispr阵列的c2c1直系同源物dic2c1和tcc2c1进行后续实验(图7a)。基因座没有携带crispr阵列使得它们的crrna:tracrrna双链体的序列不可预测。在293t细胞中共转染与靶向不同基因组位点的衍生自其他8种c2c1直系同源物的基因座的sgrna组合的dic2c1和tcc2c1以及aac2c1。t7ei测定结果表明衍生自aac2c1、akc2c1、amc2c1、bs3c2c1和lsc2c1的sgrna使tcc2c1能够稳健地编辑人类基因组(图7b、c和图8)。此外,aasgrna或aksgrna能够使tcc2c1实现多重基因组编辑(图7d和图9)。上述结果表明在来自不同系统的c2c1和双链rna之间可互换性使得可能利用天然基因座不具有crispr阵列的c2c1直系同源物来编辑哺乳动物基因组。
实施例4、设计用于c2c1介导的基因组编辑的人工sgrna
不同c2c1系统中c2c1蛋白和双rna之间的可互换性有利于设计新的人工sgrna(artsgrna)支架以促进c2c1介导的基因组编辑。考虑到c2c1直系同源物中dna序列和二级结构的保守性(图1b和3),设计并从头合成37种sgrna支架(seqidno:39-75),用于靶向人ccr5基因座(图7e,图10a)。t7ei测定的结果表明22种artsgrna支架有效地工作(图10b)。为了验证artgrna的普遍适用性,使用artsgrna13指导tcc2c1或aac2c1进行多重基因组编辑(图10a)。t7ei测定结果表明,artsgrna13同时促进tcc2c1和aac2c1两者的多重基因组编辑(图7f和图10c)。结果表明通过设计和合成artsgrna能促进c2c1介导的基因组编辑特别是多重基因组编辑。
表2本发明涉及的序列及信息
序列表
<110>中国科学院动物研究所
<120>基于c2c1核酸酶的基因组编辑系统和方法
<130>tc5170
<160>80
<170>siposequencelisting1.0
<210>1
<211>1129
<212>prt
<213>alicyclobacillusacidiphilus
<400>1
metalavallyssermetlysvallysleuargleuaspasnmetpro
151015
gluileargalaglyleutrplysleuhisthrgluvalasnalagly
202530
valargtyrtyrthrglutrpleuserleuleuargglngluasnleu
354045
tyrargargserproasnglyaspglygluglnglucystyrlysthr
505560
alagluglucyslysalagluleuleugluargleuargalaarggln
65707580
valgluasnglyhiscysglyproalaglyseraspaspgluleuleu
859095
glnleualaargglnleutyrgluleuleuvalproglnalailegly
100105110
alalysglyaspalaglnglnilealaarglyspheleuserproleu
115120125
alaasplysaspalavalglyglyleuglyilealalysalaglyasn
130135140
lysproargtrpvalargmetargglualaglygluproglytrpglu
145150155160
gluglulysalalysalaglualaarglysserthraspargthrala
165170175
aspvalleuargalaleualaasppheglyleulysproleumetarg
180185190
valtyrthraspseraspmetserservalglntrplysproleuarg
195200205
lysglyglnalavalargthrtrpaspargaspmetpheglnglnala
210215220
ilegluargmetmetsertrpglusertrpasnglnargvalglyglu
225230235240
alatyralalysleuvalgluglnlysserargphegluglnlysasn
245250255
phevalglyglngluhisleuvalglnleuvalasnglnleuglngln
260265270
aspmetlysglualaserhisglyleugluserlysgluglnthrala
275280285
histyrleuthrglyargalaleuargglyserasplysvalpheglu
290295300
lystrpglulysleuaspproaspalapropheaspleutyraspthr
305310315320
gluilelysasnvalglnargargasnthrargargpheglyserhis
325330335
aspleuphealalysleualagluprolystyrglnalaleutrparg
340345350
gluaspalaserpheleuthrargtyralavaltyrasnserileval
355360365
arglysleuasnhisalalysmetphealathrphethrleuproasp
370375380
alathralahisproiletrpthrargpheasplysleuglyglyasn
385390395400
leuhisglntyrthrpheleupheasnglupheglygluglyarghis
405410415
alaileargpheglnlysleuleuthrvalgluaspglyvalalalys
420425430
gluvalaspaspvalthrvalproilesermetseralaglnleuasp
435440445
aspleuleuproargaspprohisgluleuvalalaleutyrphegln
450455460
asptyrglyalagluglnhisleualaglyglupheglyglyalalys
465470475480
ileglntyrargargaspglnleuasnhisleuhisalaargarggly
485490495
alaargaspvaltyrleuasnleuservalargvalglnserglnser
500505510
glualaargglygluargargproprotyralaalavalpheargleu
515520525
valglyaspasnhisargalaphevalhispheasplysleuserasp
530535540
tyrleualagluhisproaspaspglylysleuglysergluglyleu
545550555560
leuserglyleuargvalmetservalaspleuglyleuargthrser
565570575
alaserileservalpheargvalalaarglysaspgluleulyspro
580585590
asnsergluglyargvalprophecyspheproilegluglyasnglu
595600605
asnleuvalalavalhisgluargserglnleuleulysleuprogly
610615620
gluthrgluserlysaspleuargalaileargglugluargglnarg
625630635640
thrleuargglnleuargthrglnleualatyrleuargleuleuval
645650655
argcysglysergluaspvalglyargarggluargsertrpalalys
660665670
leuilegluglnprometaspalaasnglnmetthrproasptrparg
675680685
glualaphegluaspgluleuglnlysleulysserleutyrglyile
690695700
cysglyaspargglutrpthrglualavaltyrgluservalargarg
705710715720
valtrparghismetglylysglnvalargasptrparglysaspval
725730735
argserglygluargprolysileargglytyrglnlysaspvalval
740745750
glyglyasnserilegluglnileglutyrleugluargglntyrlys
755760765
pheleulyssertrpserphepheglylysvalserglyglnvalile
770775780
argalaglulysglyserargphealailethrleuarggluhisile
785790795800
asphisalalysgluaspargleulyslysleualaaspargileile
805810815
metglualaleuglytyrvaltyralaleuaspaspgluargglylys
820825830
glylystrpvalalalystyrproprocysglnleuileleuleuglu
835840845
gluleuserglutyrglnpheasnasnaspargproprosergluasn
850855860
asnglnleumetglntrpserhisargglyvalpheglngluleuleu
865870875880
asnglnalaglnvalhisaspleuleuvalglythrmettyralaala
885890895
pheserserargpheaspalaargthrglyalaproglyileargcys
900905910
argargvalproalaargcysalaarggluglnasnprogluprophe
915920925
protrptrpleuasnlysphevalalagluhislysleuaspglycys
930935940
proleuargalaaspaspleuileprothrglygluglygluphephe
945950955960
valserpropheseralaglugluglyaspphehisglnilehisala
965970975
aspleuasnalaalaglnasnleuglnargargleutrpseraspphe
980985990
aspileserglnileargleuargcysasptrpglygluvalaspgly
99510001005
gluprovalleuileproargthrthrglylysargthralaaspser
101010151020
tyrglyasnlysvalphetyrthrlysthrglyvalthrtyrtyrglu
1025103010351040
arggluargglylyslysargarglysvalphealaglnglugluleu
104510501055
serglugluglualagluleuleuvalglualaaspglualaargglu
106010651070
lysservalvalleumetargaspproserglyileileasnarggly
107510801085
asptrpthrargglnlysgluphetrpsermetvalasnglnargile
109010951100
gluglytyrleuvallysglnileargserargvalargleuglnglu
1105111011151120
seralacysgluasnthrglyaspile
1125
<210>2
<211>1147
<212>prt
<213>alicyclobacilluskakegawensis
<400>2
metalavallysserilelysvallysleuargleuserglucyspro
151015
aspileleualaglymettrpglnleuhisargalathrasnalagly
202530
valargtyrtyrthrglutrpvalserleumetargglngluileleu
354045
tyrserargglyproaspglyglyglnglncystyrmetthralaglu
505560
aspcysglnarggluleuleuargargleuargasnargglnleuhis
65707580
asnglyargglnaspglnproglythraspalaaspleuleualaile
859095
serargargleutyrgluileleuvalleuglnserileglylysarg
100105110
glyaspalaglnglnilealaserserpheleuserproleuvalasp
115120125
proasnserlysglyglyargglyglualalysserglyarglyspro
130135140
alatrpglnlysmetargaspglnglyaspproargtrpvalalaala
145150155160
argglulystyrgluglnarglysalavalaspproserlysgluile
165170175
leuasnserleuaspalaleuglyleuargproleuphealavalphe
180185190
thrgluthrtyrargserglyvalasptrplysproleuglylysser
195200205
glnglyvalargthrtrpaspargaspmetpheglnglnalaleuglu
210215220
argleumetsertrpglusertrpasnargargvalglygluglutyr
225230235240
alaargleupheglnglnlysmetlysphegluglngluhispheala
245250255
gluglnserhisleuvallysleualaargalaleuglualaaspmet
260265270
argalaalaserglnglypheglualalysargglythralahisgln
275280285
ilethrargargalaleuargglyalaaspargvalphegluiletrp
290295300
lysserileprogluglualaleupheserglntyraspgluvalile
305310315320
argglnvalglnalaglulysargargasppheglyserhisaspleu
325330335
phealalysleualagluprolystyrglnproleutrpargalaasp
340345350
gluthrpheleuthrargtyralaleutyrasnglyvalleuargasp
355360365
leuglulysalaargglnphealathrphethrleuproaspalacys
370375380
valasnproiletrpthrargphegluserserglnglyserasnleu
385390395400
hislystyrglupheleupheasphisleuglyproglyarghisala
405410415
valargpheglnargleuleuvalvalglusergluglyalalysglu
420425430
argaspservalvalvalprovalalaproserglyglnleuasplys
435440445
leuvalleuargglugluglulysserservalalaleuhisleuhis
450455460
aspthralaargproaspglyphemetalaglutrpalaglyalalys
465470475480
leuglntyrgluargserthrleualaarglysalaargargasplys
485490495
glnglymetargsertrpargargglnprosermetleumetserala
500505510
alaglnmetleugluaspalalysglnalaglyaspvaltyrleuasn
515520525
ileservalargvallysserprosergluvalargglyglnargarg
530535540
proprotyralaalaleupheargileaspasplysglnargargval
545550555560
thrvalasntyrasnlysleuseralatyrleuglugluhisproasp
565570575
lysglnileproglyalaproglyleuleuserglyleuargvalmet
580585590
servalaspleuglyleuargthrseralaserileservalphearg
595600605
valalalyslysglugluvalglualaleuglyaspglyargpropro
610615620
histyrtyrproilehisglythraspaspleuvalalavalhisglu
625630635640
argserhisleuileglnmetproglygluthrgluthrlysglnleu
645650655
arglysleuargglugluargglnalavalleuargproleupheala
660665670
glnleualaleuleuargleuleuvalargcysglyalaalaaspglu
675680685
argileargthrargsertrpglnargleuthrlysglnglyargglu
690695700
phethrlysargleuthrprosertrpargglualaleugluleuglu
705710715720
leuthrargleuglualatyrcysglyargvalproaspaspglutrp
725730735
serargilevalaspargthrvalilealaleutrpargargmetgly
740745750
lysglnvalargasptrparglysglnvallysserglyalalysval
755760765
lysvallysglytyrglnleuaspvalvalglyglyasnserleuala
770775780
glnileasptyrleugluglnglntyrlyspheleuargargtrpser
785790795800
phephealaargalaserglyleuvalvalargalaasparggluser
805810815
hisphealavalalaleuargglnhisilegluasnalalysargasp
820825830
argleulyslysleualaaspargileleumetglualaleuglytyr
835840845
valtyrglualaserglyproarggluglyglntrpthralaglnhis
850855860
proprocysglnleuileileleuglugluleuseralatyrargphe
865870875880
seraspaspargproprosergluasnserlysleumetalatrpgly
885890895
hisargglyileleuglugluleuvalasnglnalaglnvalhisasp
900905910
valleuvalglythrvaltyralaalapheserserargpheaspala
915920925
argthrglyalaproglyvalargcysargargvalproalaargphe
930935940
valglyalathrvalaspaspserleuproleutrpleuthrgluphe
945950955960
leuasplyshisargleuasplysasnleuleuargproaspaspval
965970975
ileprothrglygluglyglupheleuvalserprocysglygluglu
980985990
alaalaargvalargglnvalhisalaaspileasnalaalaglnasn
99510001005
leuglnargargleutrpglnasnpheaspilethrgluleuargleu
101010151020
argcysaspvallysmetglyglygluglythrvalleuvalproarg
1025103010351040
valasnasnalaargalalysglnleupheglylyslysvalleuval
104510501055
serglnaspglyvalthrphephegluargserglnthrglyglylys
106010651070
prohisserglulysglnthraspleuthrasplysgluleugluleu
107510801085
ilealaglualaaspglualaargalalysservalvalleuphearg
109010951100
aspproserglyhisileglylysglyhistrpileargglnargglu
1105111011151120
phetrpserleuvallysglnargilegluserhisthralagluarg
112511301135
ileargvalargglyvalglyserserleuasp
11401145
<210>3
<211>1146
<212>prt
<213>alicyclobacillusmacrosporangiidus
<400>3
metasnvalalavallysserilelysvallysleumetleuglyhis
151015
leuprogluilearggluglyleutrphisleuhisglualavalasn
202530
leuglyvalargtyrtyrthrglutrpleualaleuleuargglngly
354045
asnleutyrargargglylysaspglyalaglnglucystyrmetthr
505560
alagluglncysargglngluleuleuvalargleuargasparggln
65707580
lysargasnglyhisthrglyaspproglythraspglugluleuleu
859095
glyvalalaargargleutyrgluleuleuvalproglnservalgly
100105110
lyslysglyglnalaglnmetleualaserglypheleuserproleu
115120125
alaaspprolyssergluglyglylysglythrserlysserglyarg
130135140
lysproalatrpmetglymetlysglualaglyaspserargtrpval
145150155160
glualalysalaargtyrglualaasnlysalalysaspprothrlys
165170175
glnvalilealaserleuglumettyrglyleuargproleupheasp
180185190
valphethrgluthrtyrlysthrileargtrpmetproleuglylys
195200205
hisglnglyvalargalatrpaspargaspmetpheglnglnserleu
210215220
gluargleumetsertrpglusertrpasngluargvalglyalaglu
225230235240
phealaargleuvalaspargargaspargpheargglulyshisphe
245250255
thrglyglngluhisleuvalalaleualaglnargleugluglnglu
260265270
metlysglualaserproglyphegluserlysserserglnalahis
275280285
argilethrlysargalaleuargglyalaaspglyileileaspasp
290295300
trpleulysleusergluglygluprovalaspargpheaspgluile
305310315320
leuarglysargglnalaglnasnproargargpheglyserhisasp
325330335
leupheleulysleualagluprovalpheglnproleutrpargglu
340345350
aspproserpheleuserargtrpalasertyrasngluvalleuasn
355360365
lysleugluaspalalysglnphealathrphethrleuproserpro
370375380
cysserasnprovaltrpalaargphegluasnalagluglythrasn
385390395400
ilephelystyrasppheleupheasphispheglylysglyarghis
405410415
glyvalargpheglnargmetilevalmetargaspglyvalprothr
420425430
gluvalgluglyilevalvalproilealaproserargglnleuasp
435440445
alaleualaproasnaspalaalaserproileaspvalphevalgly
450455460
aspproalaalaproglyalapheargglyglnpheglyglyalalys
465470475480
ileglntyrargargseralaleuvalarglysglyargarggluglu
485490495
lysalatyrleucysglypheargleuproserglnargargthrgly
500505510
thrproalaaspaspalaglygluvalpheleuasnleuserleuarg
515520525
valgluserglnsergluglnalaglyargargasnproprotyrala
530535540
alavalphehisileseraspglnthrargargvalilevalargtyr
545550555560
glygluilegluargtyrleualagluhisproaspthrglyilepro
565570575
glyserargglyleuthrserglyleuargvalmetservalaspleu
580585590
glyleuargthrseralaalaileservalpheargvalalahisarg
595600605
aspgluleuthrproaspalahisglyargglnprophephephepro
610615620
ilehisglymetasphisleuvalalaleuhisgluargserhisleu
625630635640
ileargleuproglygluthrgluserlyslysvalargserilearg
645650655
gluglnargleuaspargleuasnargleuargserglnmetalaser
660665670
leuargleuleuvalargthrglyvalleuaspgluglnlysargasp
675680685
argasntrpgluargleuglnsersermetgluargglyglygluarg
690695700
metproserasptrptrpaspleupheglnalaglnvalargtyrleu
705710715720
alaglnhisargaspalaserglyglualatrpglyargmetvalgln
725730735
alaalavalargthrleutrpargglnleualalysglnvalargasp
740745750
trparglysgluvalargargasnalaasplysvallysilearggly
755760765
ilealaargaspvalproglyglyhisserleualaglnleuasptyr
770775780
leugluargglntyrargpheleuargsertrpseralapheserval
785790795800
glnalaglyglnvalvalargalagluargaspserargphealaval
805810815
alaleuarggluhisileaspasnglylyslysaspargleulyslys
820825830
leualaaspargileleumetglualaleuglytyrvaltyrvalthr
835840845
aspglyargargalaglyglntrpglnalavaltyrproprocysgln
850855860
leuvalleuleuglugluleuserglutyrargpheserasnasparg
865870875880
proprosergluasnserglnleumetvaltrpserhisargglyval
885890895
leuglugluleuilehisglnalaglnvalhisaspvalleuvalgly
900905910
thrileproalaalapheserserargpheaspalaargthrglyala
915920925
proglyileargcysargargvalproserileproleulysaspala
930935940
proserileproiletrpleuserhistyrleulysglnthrgluarg
945950955960
aspalaalaalaleuargproglygluleuileprothrglyaspgly
965970975
glupheleuvalthrproalaglyargglyalaserglyvalargval
980985990
valhisalaaspileasnalaalahisasnleuglnargargleutrp
99510001005
gluasnpheaspleuseraspileargvalargcysaspargargglu
101010151020
glylysaspglythrvalvalleuileproargleuthrasnglnarg
1025103010351040
vallysgluargtyrserglyvalilephethrsergluaspglyval
104510501055
serphethrvalglyaspalalysthrargargargserseralaser
106010651070
glnglygluglyaspaspleuseraspglugluglngluleuleuala
107510801085
glualaaspaspalaarggluargservalvalleupheargasppro
109010951100
serglyphevalasnglyglyargtrpthralaglnargalaphetrp
1105111011151120
glymetvalhisasnargilegluthrleuleualagluargpheser
112511301135
valserglyalaalaglulysvalarggly
11401145
<210>4
<211>1108
<212>prt
<213>bacillushisashii
<400>4
metalathrargserpheileleulysilegluproasnglugluval
151015
lyslysglyleutrplysthrhisgluvalleuasnhisglyileala
202530
tyrtyrmetasnileleulysleuileargglnglualailetyrglu
354045
hishisgluglnaspprolysasnprolyslysvalserlysalaglu
505560
ileglnalagluleutrpaspphevalleulysmetglnlyscysasn
65707580
serphethrhisgluvalasplysaspgluvalpheasnileleuarg
859095
gluleutyrglugluleuvalproserservalglulyslysglyglu
100105110
alaasnglnleuserasnlyspheleutyrproleuvalaspproasn
115120125
serglnserglylysglythralaserserglyarglysproargtrp
130135140
tyrasnleulysilealaglyaspprosertrpglugluglulyslys
145150155160
lystrpglugluasplyslyslysaspproleualalysileleugly
165170175
lysleualaglutyrglyleuileproleupheileprotyrthrasp
180185190
serasngluproilevallysgluilelystrpmetglulysserarg
195200205
asnglnservalargargleuasplysaspmetpheileglnalaleu
210215220
gluargpheleusertrpglusertrpasnleulysvallysgluglu
225230235240
tyrglulysvalglulysglutyrlysthrleuglugluargilelys
245250255
gluaspileglnalaleulysalaleugluglntyrglulysgluarg
260265270
glngluglnleuleuargaspthrleuasnthrasnglutyrargleu
275280285
serlysargglyleuargglytrparggluileileglnlystrpleu
290295300
lysmetaspgluasngluproserglulystyrleugluvalphelys
305310315320
asptyrglnarglyshisproargglualaglyasptyrservaltyr
325330335
glupheleuserlyslysgluasnhispheiletrpargasnhispro
340345350
glutyrprotyrleutyralathrphecysgluileasplyslyslys
355360365
lysaspalalysglnglnalathrphethrleualaaspproileasn
370375380
hisproleutrpvalargpheglugluargserglyserasnleuasn
385390395400
lystyrargileleuthrgluglnleuhisthrglulysleulyslys
405410415
lysleuthrvalglnleuaspargleuiletyrprothrglusergly
420425430
glytrpgluglulysglylysvalaspilevalleuleuproserarg
435440445
glnphetyrasnglnilepheleuaspilegluglulysglylyshis
450455460
alaphethrtyrlysaspgluserilelyspheproleulysglythr
465470475480
leuglyglyalaargvalglnpheaspargasphisleuargargtyr
485490495
prohislysvalgluserglyasnvalglyargiletyrpheasnmet
500505510
thrvalasnilegluprothrgluserprovalserlysserleulys
515520525
ilehisargaspasppheprolysvalvalasnphelysprolysglu
530535540
leuthrglutrpilelysaspserlysglylyslysleulyssergly
545550555560
ilegluserleugluileglyleuargvalmetserileaspleugly
565570575
glnargglnalaalaalaalaserilephegluvalvalaspglnlys
580585590
proaspilegluglylysleuphepheproilelysglythrgluleu
595600605
tyralavalhisargalaserpheasnilelysleuproglygluthr
610615620
leuvallysserarggluvalleuarglysalaarggluaspasnleu
625630635640
lysleumetasnglnlysleuasnpheleuargasnvalleuhisphe
645650655
glnglnphegluaspilethrgluargglulysargvalthrlystrp
660665670
ileserargglngluasnseraspvalproleuvaltyrglnaspglu
675680685
leuileglnilearggluleumettyrlysprotyrlysasptrpval
690695700
alapheleulysglnleuhislysargleugluvalgluileglylys
705710715720
gluvallyshistrparglysserleuseraspglyarglysglyleu
725730735
tyrglyileserleulysasnileaspgluileaspargthrarglys
740745750
pheleuleuargtrpserleuargprothrgluproglygluvalarg
755760765
argleugluproglyglnargphealaileaspglnleuasnhisleu
770775780
asnalaleulysgluaspargleulyslysmetalaasnthrileile
785790795800
methisalaleuglytyrcystyraspvalarglyslyslystrpgln
805810815
alalysasnproalacysglnileileleuphegluaspleuserasn
820825830
tyrasnprotyrglugluargserargphegluasnserlysleumet
835840845
lystrpserargarggluileproargglnvalalaleuglnglyglu
850855860
iletyrglyleuglnvalglygluvalglyalaglnpheserserarg
865870875880
phehisalalysthrglyserproglyileargcysservalvalthr
885890895
lysglulysleuglnaspasnargphephelysasnleuglnargglu
900905910
glyargleuthrleuasplysilealavalleulysgluglyaspleu
915920925
tyrproasplysglyglyglulyspheileserleuserlysasparg
930935940
lyscysvalthrthrhisalaaspileasnalaalaglnasnleugln
945950955960
lysargphetrpthrargthrhisglyphetyrlysvaltyrcyslys
965970975
alatyrglnvalaspglyglnthrvaltyrileprogluserlysasp
980985990
glnlysglnlysileilegluglupheglygluglytyrpheileleu
99510001005
lysaspglyvaltyrglutrpvalasnalaglylysleulysilelys
101010151020
lysglyserserlysglnsersersergluleuvalaspseraspile
1025103010351040
leulysaspserpheaspleualasergluleulysglyglulysleu
104510501055
metleutyrargaspproserglyasnvalpheproserasplystrp
106010651070
metalaalaglyvalphepheglylysleugluargileleuileser
107510801085
lysleuthrasnglntyrserileserthrilegluaspaspserser
109010951100
lysglnsermet
1105
<210>5
<211>1108
<212>prt
<213>bacillus
<400>5
metalaileargserilelysleulysleulysthrhisthrglypro
151015
glualaglnasnleuarglysglyiletrpargthrhisargleuleu
202530
asngluglyvalalatyrtyrmetlysmetleuleuleuphearggln
354045
gluserthrglygluargprolysglugluleuglnglugluleuile
505560
cyshisilearggluglnglnglnargasnglnalaasplysasnthr
65707580
glnalaleuproleuasplysalaleuglualaleuargglnleutyr
859095
gluleuleuvalproserservalglyglnserglyaspalaglnile
100105110
ileserarglyspheleuserproleuvalaspproasnserglugly
115120125
glylysglythrserlysalaglyalalysprothrtrpglnlyslys
130135140
lysglualaasnaspprothrtrpgluglnasptyrglulystrplys
145150155160
lysargargglugluaspprothralaservalilethrthrleuglu
165170175
glutyrglyileargproilepheproleutyrthrasnthrvalthr
180185190
aspilealatrpleuproleuglnserasnglnphevalargthrtrp
195200205
aspargaspmetleuglnglnalailegluargleuleusertrpglu
210215220
sertrpasnlysargvalglngluglutyralalysleulysglulys
225230235240
metalaglnleuasngluglnleugluglyglyglnglutrpileser
245250255
leuleugluglntyrglugluasnarggluarggluleuarggluasn
260265270
metthralaalaasnasplystyrargilethrlysargglnmetlys
275280285
glytrpasngluleutyrgluleutrpserthrpheproalaserala
290295300
serhisgluglntyrlysglualaleulysargvalglnglnargleu
305310315320
argglyargpheglyaspalahisphepheglntyrleumetgluglu
325330335
lysasnargleuiletrplysglyasnproglnargilehistyrphe
340345350
valalaargasngluleuthrlysargleugluglualalysglnser
355360365
alathrmetthrleuproasnalaarglyshisproleutrpvalarg
370375380
pheaspalaargglyglyasnleuglnasptyrtyrleuthralaglu
385390395400
alaasplysproargserargargphevalthrpheserglnleuile
405410415
trpprosergluserglytrpmetglulyslysaspvalgluvalglu
420425430
leualaleuserargglnphetyrglnglnvallysleuleulysasn
435440445
asplysglylysglnlysilegluphelysasplysglyserglyser
450455460
thrpheasnglyhisleuglyglyalalysleuglnleugluarggly
465470475480
aspleuglulysgluglulysasnphegluaspglygluileglyser
485490495
valtyrleuasnvalvalileaspphegluproleuglngluvallys
500505510
asnglyargvalglnalaprotyrglyglnvalleuglnleuilearg
515520525
argproasnglupheprolysvalthrthrtyrlyssergluglnleu
530535540
valglutrpilelysalaserproglnhisseralaglyvalgluser
545550555560
leualaserglypheargvalmetserileaspleuglyleuargala
565570575
alaalaalathrserilepheservalglugluserserasplysasn
580585590
alaalaaspphesertyrtrpilegluglythrproleuvalalaval
595600605
hisglnargsertyrmetleuargleuproglygluglnvalglulys
610615620
glnvalmetglulysargaspgluargpheglnleuhisglnargval
625630635640
lyspheglnileargvalleualaglnilemetargmetalaasnlys
645650655
glntyrglyaspargtrpaspgluleuaspserleulysglnalaval
660665670
gluglnlyslysserproleuaspglnthraspargthrphetrpglu
675680685
glyilevalcysaspleuthrlysvalleuproargasnglualaasp
690695700
trpgluglnalavalvalglnilehisarglysalagluglutyrval
705710715720
glylysalavalglnalatrparglysargphealaalaaspgluarg
725730735
lysglyilealaglyleusermettrpasnileglugluleuglugly
740745750
leuarglysleuleuilesertrpserargargthrargasnprogln
755760765
gluvalasnargphegluargglyhisthrserhisglnargleuleu
770775780
thrhisileglnasnvallysgluaspargleulysglnleuserhis
785790795800
alailevalmetthralaleuglytyrvaltyraspgluarglysgln
805810815
glutrpcysalaglutyrproalacysglnvalileleuphegluasn
820825830
leuserglntyrargserasnleuaspargserthrlysgluasnser
835840845
thrleumetlystrpalahisargserileprolystyrvalhismet
850855860
glnalagluprotyrglyileglnileglyaspvalargalaglutyr
865870875880
serserargphetyralalysthrglythrproglyileargcyslys
885890895
lysvalargglyglnaspleuglnglyargargphegluasnleugln
900905910
lysargleuvalasngluglnpheleuthrglugluglnvallysgln
915920925
leuargproglyaspilevalproaspaspserglygluleuphemet
930935940
thrleuthraspglyserglyserlysgluvalvalpheleuglnala
945950955960
aspileasnalaalahisasnleuglnlysargphetrpglnargtyr
965970975
asngluleuphelysvalsercysargvalilevalargaspgluglu
980985990
glutyrleuvalprolysthrlysservalglnalalysleuglylys
99510001005
glyleuphevallyslysseraspthralatrplysaspvaltyrval
101010151020
trpaspserglnalalysleulysglylysthrthrphethrgluglu
1025103010351040
sergluserprogluglnleugluasppheglngluileilegluglu
104510501055
alagluglualalysglythrtyrargthrleupheargaspproser
106010651070
glyvalphepheprogluservaltrptyrproglnlysaspphetrp
107510801085
glygluvallysarglysleutyrglylysleuarggluargpheleu
109010951100
thrlysalaarg
1105
<210>6
<211>1112
<212>prt
<213>bacillus
<400>6
metalaileargserilelysleulysmetlysthrasnserglythr
151015
aspseriletyrleuarglysalaleutrpargthrhisglnleuile
202530
asngluglyilealatyrtyrmetasnleuleuthrleutyrarggln
354045
glualaileglyasplysthrlysglualatyrglnalagluleuile
505560
asnileileargasnglnglnargasnasnglyserserglugluhis
65707580
glyseraspglngluileleualaleuleuargglnleutyrgluleu
859095
ileileproserserileglygluserglyaspalaasnglnleugly
100105110
asnlyspheleutyrproleuvalaspproasnserglnserglylys
115120125
glythrserasnalaglyarglysproargtrplysargleulysglu
130135140
gluglyasnproasptrpgluleuglulyslyslysaspglugluarg
145150155160
lysalalysaspprothrvallysilepheaspasnleuasnlystyr
165170175
glyleuleuproleupheproleuphethrasnileglnlysaspile
180185190
glutrpleuproleuglylysargglnservalarglystrpasplys
195200205
aspmetpheileglnalailegluargleuleusertrpglusertrp
210215220
asnargargvalalaaspglutyrlysglnleulysglulysthrglu
225230235240
sertyrtyrlysgluhisleuthrglyglygluglutrpileglulys
245250255
ilearglyspheglulysgluargasnmetgluleuglulysasnala
260265270
phealaproasnaspglytyrpheilethrserargglnilearggly
275280285
trpaspargvaltyrglulystrpserlysleuprogluseralaser
290295300
proglugluleutrplysvalvalalagluglnglnasnlysmetser
305310315320
gluglypheglyaspprolysvalpheserpheleualaasnargglu
325330335
asnargaspiletrpargglyhissergluargiletyrhisileala
340345350
alatyrasnglyleuglnlyslysleuserargthrlysgluglnala
355360365
thrphethrleuproaspalailegluhisproleutrpileargtyr
370375380
gluserproglyglythrasnleuasnleuphelysleugluglulys
385390395400
glnlyslysasntyrtyrvalthrleuserlysileiletrpproser
405410415
gluglulystrpileglulysgluasnilegluileproleualapro
420425430
serileglnpheasnargglnilelysleulysglnhisvallysgly
435440445
lysglngluileserpheserasptyrserserargileserleuasp
450455460
glyvalleuglyglyserargileglnpheasnarglystyrilelys
465470475480
asnhislysgluleuleuglygluglyaspileglyprovalphephe
485490495
asnleuvalvalaspvalalaproleuglngluthrargasnglyarg
500505510
leuglnserproileglylysalaleulysvalileserseraspphe
515520525
serlysvalileasptyrlysprolysgluleumetasptrpmetasn
530535540
thrglyseralaserasnserpheglyvalalaserleuleuglugly
545550555560
metargvalmetserileaspmetglyglnargthrseralaserval
565570575
serilephegluvalvallysgluleuprolysaspglngluglnlys
580585590
leuphetyrserileasnaspthrgluleuphealailehislysarg
595600605
serpheleuleuasnleuproglygluvalvalthrlysasnasnlys
610615620
glnglnargglngluargarglyslysargglnphevalargsergln
625630635640
ileargmetleualaasnvalleuargleugluthrlyslysthrpro
645650655
aspgluarglyslysalailehislysleumetgluilevalglnser
660665670
tyraspsertrpthralaserglnlysgluvaltrpglulysgluleu
675680685
asnleuleuthrasnmetalaalapheasnaspgluiletrplysglu
690695700
serleuvalgluleuhishisargilegluprotyrvalglyglnile
705710715720
valserlystrparglysglyleusergluglyarglysasnleuala
725730735
glyilesermettrpasnileaspgluleugluaspthrargargleu
740745750
leuilesertrpserlysargserargthrproglyglualaasnarg
755760765
ilegluthraspglupropheglyserserleuleuglnhisilegln
770775780
asnvallysaspaspargleulysglnmetalaasnleuileilemet
785790795800
thralaleuglyphelystyrasplysgluglulysaspargtyrlys
805810815
argtrplysgluthrtyrproalacysglnileileleuphegluasn
820825830
leuasnargtyrleupheasnleuaspargserargarggluasnser
835840845
argleumetlystrpalahisargserileproargthrvalsermet
850855860
glnglyglumetpheglyleuglnvalglyaspvalargserglutyr
865870875880
serserargphehisalalysthrglyalaproglyileargcyshis
885890895
alaleuthrglugluaspleulysalaglyserasnthrleulysarg
900905910
leuilegluaspglypheileasnglusergluleualatyrleulys
915920925
lysglyaspileileproserglnglyglygluleuphevalthrleu
930935940
serlysargtyrlyslysaspseraspasnasngluleuthrvalile
945950955960
hisalaaspileasnalaalaglnasnleuglnlysargphetrpgln
965970975
glnasnsergluvaltyrargvalprocysglnleualaargmetgly
980985990
gluasplysleutyrileprolysserglnthrgluthrilelyslys
99510001005
tyrpheglylysglyserphevallysasnasnthrgluglngluval
101010151020
tyrlystrpglulysserglulysmetlysilelysthraspthrthr
1025103010351040
pheaspleuglnaspleuaspglyphegluaspileserlysthrile
104510501055
gluleualaglngluglnglnlyslystyrleuthrmetpheargasp
106010651070
proserglytyrphepheasnasngluthrtrpargproglnlysglu
107510801085
tyrtrpserilevalasnasnileilelyssercysleulyslyslys
109010951100
ileleuserasnlysvalgluleu
11051110
<210>7
<211>1149
<212>prt
<213>desulfovibrioinopinatus
<400>7
metprothrargthrileasnleulysleuvalleuglylysasnpro
151015
gluasnalathrleuargargalaleupheserthrhisargleuval
202530
asnglnalathrlysargilegluglupheleuleuleucysarggly
354045
glualatyrargthrvalaspasngluglylysglualagluilepro
505560
arghisalavalglngluglualaleualaphealalysalaalagln
65707580
arghisasnglycysileserthrtyrgluaspglngluileleuasp
859095
valleuargglnleutyrgluargleuvalproservalasngluasn
100105110
asnglualaglyaspalaglnalaalaasnalatrpvalserproleu
115120125
metseralaglusergluglyglyleuservaltyrasplysvalleu
130135140
aspproproprovaltrpmetlysleulysgluglulysalaprogly
145150155160
trpglualaalaserglniletrpileglnseraspgluglyglnser
165170175
leuleuasnlysproglyserproproargtrpilearglysleuarg
180185190
serglyglnprotrpglnaspaspphevalseraspglnlyslyslys
195200205
glnaspgluleuthrlysglyasnalaproleuilelysglnleulys
210215220
glumetglyleuleuproleuvalasnprophephearghisleuleu
225230235240
aspprogluglylysglyvalserprotrpaspargleualavalarg
245250255
alaalavalalahispheilesertrpglusertrpasnhisargthr
260265270
argalaglutyrasnserleulysleuargargaspgluphegluala
275280285
alaseraspgluphelysaspaspphethrleuleuargglntyrglu
290295300
alalysarghisserthrleulysserilealaleualaaspaspser
305310315320
asnprotyrargileglyvalargserleuargalatrpasnargval
325330335
arggluglutrpileasplysglyalathrglugluglnargvalthr
340345350
ileleuserlysleuglnthrglnleuargglylyspheglyasppro
355360365
aspleupheasntrpleualaglnasparghisvalhisleutrpser
370375380
proargaspservalthrproleuvalargileasnalavalasplys
385390395400
valleuargargarglysprotyralaleumetthrphealahispro
405410415
argphehisproargtrpileleutyrglualaproglyglyserasn
420425430
leuargglntyralaleuaspcysthrgluasnalaleuhisilethr
435440445
leuproleuleuvalaspaspalahisglythrtrpileglulyslys
450455460
ileargvalproleualaproserglyglnileglnaspleuthrleu
465470475480
glulysleuglulyslyslysasnargleutyrtyrargserglyphe
485490495
glnglnphealaglyleualaglyglyalagluvalleuphehisarg
500505510
protyrmetgluhisaspgluargserglugluserleuleugluarg
515520525
proglyalavaltrpphelysleuthrleuaspvalalathrglnala
530535540
proproasntrpleuaspglylysglyargvalargthrproproglu
545550555560
valhishisphelysthralaleuserasnlysserlyshisthrarg
565570575
thrleuglnproglyleuargvalleuservalaspleuglymetarg
580585590
thrphealasercysservalphegluleuilegluglylysproglu
595600605
thrglyargalapheprovalalaaspgluargsermetaspserpro
610615620
asnlysleutrpalalyshisgluargserphelysleuthrleupro
625630635640
glygluthrproserarglysglugluglugluargserilealaarg
645650655
alagluiletyralaleulysargaspileglnargleulysserleu
660665670
leuargleuglyglugluaspasnaspasnargargaspalaleuleu
675680685
gluglnphephelysglytrpglyglugluaspvalvalproglygln
690695700
alapheproargserleupheglnglyleuglyalaalaprophearg
705710715720
serthrprogluleutrpargglnhiscysglnthrtyrtyrasplys
725730735
alaglualacysleualalyshisileserasptrparglysargthr
740745750
argproargprothrserargglumettrptyrlysthrargsertyr
755760765
hisglyglylysseriletrpmetleuglutyrleuaspalavalarg
770775780
lysleuleuleusertrpserleuargglyargthrtyrglyalaile
785790795800
asnargglnaspthralaargpheglyserleualaserargleuleu
805810815
hishisileasnserleulysgluaspargilelysthrglyalaasp
820825830
serilevalglnalaalaargglytyrileproleuprohisglylys
835840845
glytrpgluglnargtyrgluprocysglnleuileleuphegluasp
850855860
leualaargtyrargpheargvalaspargproargarggluasnser
865870875880
glnleumetglntrpasnhisargalailevalalagluthrthrmet
885890895
glnalagluleutyrglyglnilevalgluasnthralaalaglyphe
900905910
serserargphehisalaalathrglyalaproglyvalargcysarg
915920925
pheleuleugluargasppheaspasnaspleuprolysprotyrleu
930935940
leuarggluleusertrpmetleuglyasnthrlysvalgluserglu
945950955960
gluglulysleuargleuleuserglulysileargproglyserleu
965970975
valprotrpaspglyglygluglnphealathrleuhisprolysarg
980985990
glnthrleucysvalilehisalaaspmetasnalaalaglnasnleu
99510001005
glnargargphepheglyargcysglyglualapheargleuvalcys
101010151020
glnprohisglyaspaspvalleuargleualaserthrproglyala
1025103010351040
argleuleuglyalaleuglnglnleugluasnglyglnglyalaphe
104510501055
gluleuvalargaspmetglyserthrserglnmetasnargpheval
106010651070
metlysserleuglylyslyslysilelysproleuglnaspasnasn
107510801085
glyaspaspgluleugluaspvalleuservalleuproglugluasp
109010951100
aspthrglyargilethrvalpheargaspserserglyilephephe
1105111011151120
procysasnvaltrpileproalalysglnphetrpproalavalarg
112511301135
alametiletrplysvalmetalaserhisserleugly
11401145
<210>8
<211>1090
<212>prt
<213>laceyellasediminis
<400>8
metserileargserphelysleulysilelysthrlysserglyval
151015
asnalaglugluleuargargglyleutrpargthrhisglnleuile
202530
asnaspglyilealatyrtyrmetasntrpleuvalleuleuarggln
354045
gluaspleupheileargasnglugluthrasngluileglulysarg
505560
serlysglugluileglnglygluleuleugluargvalhislysgln
65707580
glnglnargasnglntrpserglygluvalaspaspglnthrleuleu
859095
glnthrleuarghisleutyrglugluilevalproservalilegly
100105110
lysserglyasnalaserleulysalaargphepheleuglyproleu
115120125
valaspproasnasnlysthrthrlysaspvalserlysserglypro
130135140
thrprolystrplyslysmetlysaspalaglyaspproasntrpval
145150155160
glnglutyrglulystyrmetalagluargglnthrleuvalargleu
165170175
gluglumetglyleuileproleuphepromettyrthraspgluval
180185190
glyaspilehistrpleuproglnalaserglytyrthrargthrtrp
195200205
aspargaspmetpheglnglnalailegluargleuleusertrpglu
210215220
sertrpasnargargvalarggluargargalaglnpheglulyslys
225230235240
thrhisaspphealaserargphesergluseraspvalglntrpmet
245250255
asnlysleuargglutyrglualaglnglnglulysserleugluglu
260265270
asnalaphealaproasngluprotyralaleuthrlyslysalaleu
275280285
argglytrpgluargvaltyrhissertrpmetargleuaspserala
290295300
alasergluglualatyrtrpglngluvalalathrcysglnthrala
305310315320
metargglyglupheglyaspproalailetyrglnpheleualagln
325330335
lysgluasnhisaspiletrpargglytyrprogluargvalileasp
340345350
phealagluleuasnhisleuglnarggluleuargargalalysglu
355360365
aspalathrphethrleuproaspservalasphisproleutrpval
370375380
argtyrglualaproglyglythrasnilehisglytyraspleuval
385390395400
glnaspthrlysargasnleuthrleuileleuasplyspheileleu
405410415
proaspgluasnglysertrphisgluvallyslysvalpropheser
420425430
leualalysserlysglnphehisargglnvaltrpleuglngluglu
435440445
glnlysglnlyslysarggluvalvalphetyrasptyrserthrasn
450455460
leuprohisleuglythrleualaglyalalysleuglntrpasparg
465470475480
asnpheleuasnlysargthrglnglnglnileglugluthrglyglu
485490495
ileglylysvalphepheasnileservalaspvalargproalaval
500505510
gluvallysasnglyargleuglnasnglyleuglylysalaleuthr
515520525
valleuthrhisproaspglythrlysilevalthrglytrplysala
530535540
gluglnleuglulystrpvalglygluserglyargvalserserleu
545550555560
glyleuaspserleusergluglyleuargvalmetserileaspleu
565570575
glyglnargthrseralathrvalservalphegluilethrlysglu
580585590
alaproaspasnprotyrlysphephetyrglnleugluglythrglu
595600605
leuphealavalhisglnargserpheleuleualaleuproglyglu
610615620
asnproproglnlysilelysglnmetarggluileargtrplysglu
625630635640
argasnargilelysglnglnvalaspglnleuseralaileleuarg
645650655
leuhislyslysvalasngluaspgluargileglnalaileasplys
660665670
leuleuglnlysvalalasertrpglnleuasnglugluilealathr
675680685
alatrpasnglnalaleuserglnleutyrserlysalalysgluasn
690695700
aspleuglntrpasnglnalailelysasnalahishisglnleuglu
705710715720
provalvalglylysglnileserleutrparglysaspleuserthr
725730735
glyargglnglyilealaglyleuserleutrpserileglugluleu
740745750
glualathrlyslysleuleuthrargtrpserlysargserargglu
755760765
proglyvalvallysargilegluargphegluthrphealalysgln
770775780
ileglnhishisileasnglnvallysgluasnargleulysglnleu
785790795800
alaasnleuilevalmetthralaleuglytyrlystyraspglnglu
805810815
glnlyslystrpilegluvaltyrproalacysglnvalvalleuphe
820825830
gluasnleuargsertyrargphesertyrgluargserargargglu
835840845
asnlyslysleumetglutrpserhisargserileprolysleuval
850855860
glnmetglnglygluleupheglyleuglnvalalaaspvaltyrala
865870875880
alatyrserserargtyrhisglyargthrglyalaproglyilearg
885890895
cyshisalaleuthrglualaaspleuargasngluthrasnileile
900905910
hisgluleuileglualaglypheilelysglugluhisargprotyr
915920925
leuglnglnglyaspleuvalprotrpserglyglygluleupheala
930935940
thrleuglnlysprotyraspasnproargileleuthrleuhisala
945950955960
aspileasnalaalaglnasnileglnlysargphetrphisproser
965970975
mettrppheargvalasncysgluservalmetgluglygluileval
980985990
thrtyrvalprolysasnlysthrvalhislyslysglnglylysthr
99510001005
pheargphevallysvalgluglyseraspvaltyrglutrpalalys
101010151020
trpserlysasnargasnlysasnthrpheserserilethrgluarg
1025103010351040
lysproprosersermetileleupheargaspproserglythrphe
104510501055
phelysgluglnglutrpvalgluglnlysthrphetrpglylysval
106010651070
glnsermetileglnalatyrmetlyslysthrilevalglnargmet
107510801085
gluglu
1090
<210>9
<211>1119
<212>prt
<213>spirochaetes
<400>9
metserphethrilesertyrprophelysleuileilelysasnlys
151015
aspglualalysalaleuleuaspthrhisglntyrmetasnglugly
202530
vallystyrtyrleuglulysleuleumetpheargglnglulysile
354045
pheileglygluaspgluthrglylysargiletyrileglugluthr
505560
glutyrlyslysglnileglugluphetyrleuilelyslysthrglu
65707580
leuglyargasnleuthrleuthrleuaspgluphelysthrleumet
859095
arggluleutyrilecysleuvalsersersermetgluasnlyslys
100105110
glypheproasnalaglnglnalaserleuasnilepheserproleu
115120125
pheaspalagluserlysglytyrileleulysglugluasnasnasn
130135140
ileserleuilehislysasptyrglylysileleuleulysargleu
145150155160
argaspasnasnleuileproilephethrlysphethraspilelys
165170175
lysilethralalysleuserprothralaleuaspargmetilephe
180185190
alaglnalaileglulysleuleusertyrglusertrpcyslysleu
195200205
metilelysgluargpheasplysgluvallysilelysgluleuglu
210215220
asnlyscysgluasnlysglngluargasplysilephegluileleu
225230235240
glulystyrgluglugluargglnlysthrphegluglnaspsergly
245250255
phealalyslysglylysphetyrilethrglyargmetleulysgly
260265270
pheaspgluilelysglulystrpleulysglulysaspargserglu
275280285
glnasnleuileasnileleuasnlystyrglnthraspasnserlys
290295300
leuvalglyaspargasnleupheglupheileilelysleugluasn
305310315320
glncysleutrpasnglyaspileasptyrleulysilelysargasp
325330335
ileasnlysasnglniletrpleuaspargproglumetproargphe
340345350
thrmetproaspphelyslyshisproleutrptyrargtyrgluasp
355360365
proserasnserasnpheargasntyrlysilegluvalvallysasp
370375380
gluasntyrilethrileproleuilethrgluargasnasnglutyr
385390395400
pheglugluasntyrthrpheasnleualalysleulyslysleuser
405410415
gluasnilethrpheileprolysserlysasnlysgluphegluphe
420425430
ileaspserasnaspgluglugluasplyslysaspglnlyslysser
435440445
lysglntyrilelystyrcysaspthralalysasnthrsertyrgly
450455460
lysserglyglyileargleutyrpheasnargasngluleugluasn
465470475480
tyrlysaspglylyslysmetaspsertyrthrvalphethrleuser
485490495
ileargasptyrlysserleuphealalysglulysleuglnprogln
500505510
ilepheasnthrvalaspasnlysilethrserleulysileglnlys
515520525
lyspheglyasnglugluglnthrasnpheleusertyrphethrgln
530535540
asnglnilethrlyslysasptrpmetaspglulysthrpheglnasn
545550555560
vallysgluleuasngluglyileargvalleuservalaspleugly
565570575
glnargphephealaalavalsercysphegluilemetsergluile
580585590
aspasnasnlysleuphepheasnleuasnaspglnasnhislysile
595600605
ileargileasnasplysasntyrtyralalyshisiletyrserlys
610615620
thrilelysleuserglygluaspaspaspleutyrlysgluarglys
625630635640
ileasnlysasntyrlysleusertyrglngluarglysasnlysile
645650655
glyilephethrargglnileasnlysleuasnglnleuleulysile
660665670
ileargasnaspgluileasplysglulysphelysgluleuileglu
675680685
thrthrlysargtyrvallysasnthrtyrasnaspglyileileasp
690695700
trpasnasnvalaspasnlysileleusertyrgluasnlysgluasp
705710715720
valileasnleuhislysgluleuasplyslysleugluileaspphe
725730735
lysglupheileargglucysarglysproilepheargserglygly
740745750
leusermetglnargileasppheleuglulysleuasnlysleulys
755760765
arglystrpvalalaargthrglnlysseralagluserilevalleu
770775780
thrprolyspheglytyrlysleulysgluhisileasngluleulys
785790795800
aspasnargvallysglnglyvalasntyrileleumetthralaleu
805810815
glytyrilelysaspasngluilelysasnaspserlyslyslysgln
820825830
lysgluasptrpvallyslysasnargalacysglnileileleumet
835840845
glulysleuthrglutyrthrphealagluaspargproarggluglu
850855860
asnserlysleuargmettrpserhisargglnilepheasnpheleu
865870875880
glnglnlysalaserleutrpglyileleuvalglyaspvalpheala
885890895
protyrthrserlyscysleuseraspasnasnalaproglyilearg
900905910
cyshisglnvalthrlyslysaspleuileaspasnsertrppheleu
915920925
lysilevalvallysaspaspalaphecysaspleuilegluileasn
930935940
lysgluasnvallysasnlysserilelysileasnaspileleupro
945950955960
leuargglyglygluleuphealaserilelysaspglylysleuhis
965970975
ilevalglnalaaspileasnalaserargasnilealalysargphe
980985990
leuserglnileasnpropheargvalvalleulyslysasplysasp
99510001005
gluthrphehisleulysasngluproasntyrleulysasntyrtyr
101010151020
serileleuasnphevalprothrasnglugluleuthrphephelys
1025103010351040
valglugluasnlysaspilelysprothrlysargilelysmetasp
104510501055
lyshisglulysgluserthraspgluglyaspasptyrserlysasn
106010651070
glnilealaleupheargaspaspserglyilephepheasplysser
107510801085
leutrpvalaspglylysilephetrpservalvallysasnlysmet
109010951100
thrlysleuleuarggluargasnasnlyslysasnglyserlys
110511101115
<210>10
<211>1142
<212>prt
<213>tuberibacilluscalidus
<400>10
metasnilehisleulysgluleuileargmetalathrlysserphe
151015
ileleulysmetlysthrlysasnasnproglnleuargleuserleu
202530
trplysthrhisgluleupheasnpheglyvalalatyrtyrmetasp
354045
leuleuserleupheargglnlysaspleutyrmethisasnaspglu
505560
aspproasphisprovalvalleulyslysglugluileglngluarg
65707580
leutrpmetlysvalarggluthrglnglnlysasnglyphehisgly
859095
gluvalserlysaspgluvalleugluthrleuargalaleutyrglu
100105110
gluleuvalproseralavalglylysserglyglualaasnglnile
115120125
serasnlystyrleutyrproleuthraspproalaserglnsergly
130135140
lysglythralaasnserglyarglysproargtrplyslysleulys
145150155160
glualaglyaspprosertrplysaspalatyrglulystrpglulys
165170175
gluargglngluaspprolysleulysileleualaalaleuglnser
180185190
pheglyleuileproleupheargprophethrgluasnasphislys
195200205
alavalileservallystrpmetprolysserlysasnglnserval
210215220
arglyspheasplysaspmetpheasnglnalailegluargpheleu
225230235240
sertrpglusertrpasnglulysvalalagluasptyrglulysthr
245250255
valseriletyrgluserleuglnlysgluleulysglyileserthr
260265270
lysalaphegluilemetgluargvalglulysalatyrglualahis
275280285
leuarggluilethrpheserasnserthrtyrargileglyasnarg
290295300
alaileargglytrpthrgluilevallyslystrpmetlysleuasp
305310315320
proseralaproglnglyasntyrleuaspvalvallysasptyrgln
325330335
argarghisproarggluserglyaspphelysleuphegluleuleu
340345350
serargprogluasnglnalaalatrpargglutyrproglupheleu
355360365
proleutyrvallystyrarghisalagluglnargmetlysthrala
370375380
lyslysglnalathrphethrleucysaspproilearghisproleu
385390395400
trpvalargtyrglugluargserglythrasnleuasnlystyrarg
405410415
leuilemetasnglulysglulysvalvalglnpheaspargleuile
420425430
cysleuasnalaaspglyhistyrglugluglngluaspvalthrval
435440445
proleualaproserglnglnpheaspaspglnilelyspheserser
450455460
gluaspthrglylysglylyshisasnphesertyrtyrhislysgly
465470475480
ileasntyrgluleulysglythrleuglyglyalaargileglnphe
485490495
asparggluhisleuleuargargglnglyvallysalaglyasnval
500505510
glyargilepheleuasnvalthrleuasnilegluprometglnpro
515520525
pheserargserglyasnleuglnthrservalglylysalaleulys
530535540
valtyrvalaspglytyrprolysvalvalasnphelysprolysglu
545550555560
leuthrgluhisilelysgluserglulysasnthrleuthrleugly
565570575
valgluserleuprothrglyleuargvalmetservalaspleugly
580585590
glnargglnalaalaalaileserilephegluvalvalserglulys
595600605
proaspaspasnlysleuphetyrprovallysaspthraspleuphe
610615620
alavalhisargthrserpheasnilelysleuproglyglulysarg
625630635640
thrgluargargmetleugluglnglnlysargaspglnalailearg
645650655
aspleuserarglysleulyspheleulysasnvalleuasnmetgln
660665670
lysleuglulysthraspgluargglulysargvalasnargtrpile
675680685
lysasparggluarggluglugluasnprovaltyrvalglngluphe
690695700
glumetileserlysvalleutyrserprohisservaltrpvalasp
705710715720
glnleulysserilehisarglysleuglugluglnleuglylysglu
725730735
ileserlystrpargglnserileserglnglyargglnglyvaltyr
740745750
glyileserleulysasnilegluaspileglulysthrargargleu
755760765
leupheargtrpsermetargprogluasnproglygluvallysgln
770775780
leuglnproglygluargphealaileaspglnglnasnhisleuasn
785790795800
hisleulysaspaspargilelyslysleualaasnglnilevalmet
805810815
thralaleuglytyrargtyraspglylysarglyslystrpileala
820825830
lyshisproalacysglnleuvalleuphegluaspleuserargtyr
835840845
alaphetyraspgluargserargleugluasnargasnleumetarg
850855860
trpserargarggluileprolysglnvalalaglnileglyglyleu
865870875880
tyrglyleuleuvalglygluvalglyalaglntyrserserargphe
885890895
hisalalysserglyalaproglyileargcysargvalvallysglu
900905910
hisgluleutyrilethrgluglyglyglnlysvalargasnglnlys
915920925
pheleuaspserleuvalgluasnasnileilegluproaspaspala
930935940
argargleugluproglyaspleuileargaspglnglyglyasplys
945950955960
phealathrleuaspgluargglygluleuvalilethrhisalaasp
965970975
ileasnalaalaglnasnleuglnlysargphetrpthrargthrhis
980985990
glyleutyrargileargcysgluserarggluilelysaspalaval
99510001005
valleuvalproserasplysaspglnlysglulysmetgluasnleu
101010151020
pheglyileglytyrleuglnprophelysglngluasnaspvaltyr
1025103010351040
lystrpvallysglyglulysilelysglylyslysthrsersergln
104510501055
seraspasplysgluleuvalsergluileleuglnglualaserval
106010651070
metalaaspgluleulysglyasnarglysthrleupheargasppro
107510801085
serglytyrvalpheprolysaspargtrptyrthrglyglyargtyr
109010951100
pheglythrleugluhisleuleulysarglysleualagluargarg
1105111011151120
leupheaspglyglyserserargargglyleupheasnglythrasp
112511301135
serasnthrasnvalglu
1140
<210>11
<211>3387
<212>dna
<213>alicyclobacillusacidiphilus
<400>11
atggccgtgaagagcatgaaggtgaagctgcgcctggacaacatgcccgagatccgcgcc60
ggcctgtggaagctgcacaccgaggtgaacgccggcgtgcgctactacaccgagtggctg120
agcctgctgcgccaggagaacctgtaccgccgcagccccaacggcgacggcgagcaggag180
tgctacaagaccgccgaggagtgcaaggccgagctgctggagcgcctgcgcgcccgccag240
gtggagaacggccactgcggccccgccggcagcgacgacgagctgctgcagctggcccgc300
cagctgtacgagctgctggtgccccaggccatcggcgccaagggcgacgcccagcagatc360
gcccgcaagttcctgagccccctggccgacaaggacgccgtgggcggcctgggcatcgcc420
aaggccggcaacaagccccgctgggtgcgcatgcgcgaggccggcgagcccggctgggag480
gaggagaaggccaaggccgaggcccgcaagagcaccgaccgcaccgccgacgtgctgcgc540
gccctggccgacttcggcctgaagcccctgatgcgcgtgtacaccgacagcgacatgagc600
agcgtgcagtggaagcccctgcgcaagggccaggccgtgcgcacctgggaccgcgacatg660
ttccagcaggccatcgagcgcatgatgagctgggagagctggaaccagcgcgtgggcgag720
gcctacgccaagctggtggagcagaagagccgcttcgagcagaagaacttcgtgggccag780
gagcacctggtgcagctggtgaaccagctgcagcaggacatgaaggaggccagccacggc840
ctggagagcaaggagcagaccgcccactacctgaccggccgcgccctgcgcggcagcgac900
aaggtgttcgagaagtgggagaagctggaccccgacgcccccttcgacctgtacgacacc960
gagatcaagaacgtgcagcgccgcaacacccgccgcttcggcagccacgacctgttcgcc1020
aagctggccgagcccaagtaccaggccctgtggcgcgaggacgccagcttcctgacccgc1080
tacgccgtgtacaacagcatcgtgcgcaagctgaaccacgccaagatgttcgccaccttc1140
accctgcccgacgccaccgcccaccccatctggacccgcttcgacaagctgggcggcaac1200
ctgcaccagtacaccttcctgttcaacgagttcggcgagggccgccacgccatccgcttc1260
cagaagctgctgaccgtggaggacggcgtggccaaggaggtggacgacgtgaccgtgccc1320
atcagcatgagcgcccagctggacgacctgctgccccgcgacccccacgagctggtggcc1380
ctgtacttccaggactacggcgccgagcagcacctggccggcgagttcggcggcgccaag1440
atccagtaccgccgcgaccagctgaaccacctgcacgcccgccgcggcgcccgcgacgtg1500
tacctgaacctgagcgtgcgcgtgcagagccagagcgaggcccgcggcgagcgccgcccc1560
ccctacgccgccgtgttccgcctggtgggcgacaaccaccgcgccttcgtgcacttcgac1620
aagctgagcgactacctggccgagcaccccgacgacggcaagctgggcagcgagggcctg1680
ctgagcggcctgcgcgtgatgagcgtggacctgggcctgcgcaccagcgccagcatcagc1740
gtgttccgcgtggcccgcaaggacgagctgaagcccaacagcgagggccgcgtgcccttc1800
tgcttccccatcgagggcaacgagaacctggtggccgtgcacgagcgcagccagctgctg1860
aagctgcccggcgagaccgagagcaaggacctgcgcgccatccgcgaggagcgccagcgc1920
accctgcgccagctgcgcacccagctggcctacctgcgcctgctggtgcgctgcggcagc1980
gaggacgtgggccgccgcgagcgcagctgggccaagctgatcgagcagcccatggacgcc2040
aaccagatgacccccgactggcgcgaggccttcgaggacgagctgcagaagctgaagagc2100
ctgtacggcatctgcggcgaccgcgagtggaccgaggccgtgtacgagagcgtgcgccgc2160
gtgtggcgccacatgggcaagcaggtgcgcgactggcgcaaggacgtgcgcagcggcgag2220
cgccccaagatccgcggctaccagaaggacgtggtgggcggcaacagcatcgagcagatc2280
gagtacctggagcgccagtacaagttcctgaagagctggagcttcttcggcaaggtgagc2340
ggccaggtgatccgcgccgagaagggcagccgcttcgccatcaccctgcgcgagcacatc2400
gaccacgccaaggaggaccgcctgaagaagctggccgaccgcatcatcatggaggccctg2460
ggctacgtgtacgccctggacgacgagcgcggcaagggcaagtgggtggccaagtacccc2520
ccctgccagctgatcctgctggaggagctgagcgagtaccagttcaacaacgaccgcccc2580
cccagcgagaacaaccagctgatgcagtggagccaccgcggcgtgttccaggagctgctg2640
aaccaggcccaggtgcacgacctgctggtgggcaccatgtacgccgccttcagcagccgc2700
ttcgacgcccgcaccggcgcccccggcatccgctgccgccgcgtgcccgcccgctgcgcc2760
cgcgagcagaaccccgagcccttcccctggtggctgaacaagttcgtggccgagcacaag2820
ctggacggctgccccctgcgcgccgacgacctgatccccaccggcgagggcgagttcttc2880
gtgagccccttcagcgccgaggagggcgacttccaccagatccacgccgacctgaacgcc2940
gcccagaacctgcagcgccgcctgtggagcgacttcgacatcagccagatccgcctgcgc3000
tgcgactggggcgaggtggacggcgagcccgtgctgatcccccgcaccaccggcaagcgc3060
accgccgacagctacggcaacaaggtgttctacaccaagaccggcgtgacctactacgag3120
cgcgagcgcggcaagaagcgccgcaaggtgttcgcccaggaggagctgagcgaggaggag3180
gccgagctgctggtggaggccgacgaggcccgcgagaagagcgtggtgctgatgcgcgac3240
cccagcggcatcatcaaccgcggcgactggacccgccagaaggagttctggagcatggtg3300
aaccagcgcatcgagggctacctggtgaagcagatccgcagccgcgtgcgcctgcaggag3360
agcgcctgcgagaacaccggcgacatc3387
<210>12
<211>3441
<212>dna
<213>alicyclobacilluskakegawensis
<400>12
atggccgtgaagagcatcaaggtgaagctgcgcctgagcgagtgccccgacatcctggcc60
ggcatgtggcagctgcaccgcgccaccaacgccggcgtgcgctactacaccgagtgggtg120
agcctgatgcgccaggagatcctgtacagccgcggccccgacggcggccagcagtgctac180
atgaccgccgaggactgccagcgcgagctgctgcgccgcctgcgcaaccgccagctgcac240
aacggccgccaggaccagcccggcaccgacgccgacctgctggccatcagccgccgcctg300
tacgagatcctggtgctgcagagcatcggcaagcgcggcgacgcccagcagatcgccagc360
agcttcctgagccccctggtggaccccaacagcaagggcggccgcggcgaggccaagagc420
ggccgcaagcccgcctggcagaagatgcgcgaccagggcgacccccgctgggtggccgcc480
cgcgagaagtacgagcagcgcaaggccgtggaccccagcaaggagatcctgaacagcctg540
gacgccctgggcctgcgccccctgttcgccgtgttcaccgagacctaccgcagcggcgtg600
gactggaagcccctgggcaagagccagggcgtgcgcacctgggaccgcgacatgttccag660
caggccctggagcgcctgatgagctgggagagctggaaccgccgcgtgggcgaggagtac720
gcccgcctgttccagcagaagatgaagttcgagcaggagcacttcgccgagcagagccac780
ctggtgaagctggcccgcgccctggaggccgacatgcgcgccgccagccagggcttcgag840
gccaagcgcggcaccgcccaccagatcacccgccgcgccctgcgcggcgccgaccgcgtg900
ttcgagatctggaagagcatccccgaggaggccctgttcagccagtacgacgaggtgatc960
cgccaggtgcaggccgagaagcgccgcgacttcggcagccacgacctgttcgccaagctg1020
gccgagcccaagtaccagcccctgtggcgcgccgacgagaccttcctgacccgctacgcc1080
ctgtacaacggcgtgctgcgcgacctggagaaggcccgccagttcgccaccttcaccctg1140
cccgacgcctgcgtgaaccccatctggacccgcttcgagagcagccagggcagcaacctg1200
cacaagtacgagttcctgttcgaccacctgggccccggccgccacgccgtgcgcttccag1260
cgcctgctggtggtggagagcgagggcgccaaggagcgcgacagcgtggtggtgcccgtg1320
gcccccagcggccagctggacaagctggtgctgcgcgaggaggagaagagcagcgtggcc1380
ctgcacctgcacgacaccgcccgccccgacggcttcatggccgagtgggccggcgccaag1440
ctgcagtacgagcgcagcaccctggcccgcaaggcccgccgcgacaagcagggcatgcgc1500
agctggcgccgccagcccagcatgctgatgagcgccgcccagatgctggaggacgccaag1560
caggccggcgacgtgtacctgaacatcagcgtgcgcgtgaagagccccagcgaggtgcgc1620
ggccagcgccgccccccctacgccgccctgttccgcatcgacgacaagcagcgccgcgtg1680
accgtgaactacaacaagctgagcgcctacctggaggagcaccccgacaagcagatcccc1740
ggcgcccccggcctgctgagcggcctgcgcgtgatgagcgtggacctgggcctgcgcacc1800
agcgccagcatcagcgtgttccgcgtggccaagaaggaggaggtggaggccctgggcgac1860
ggccgccccccccactactaccccatccacggcaccgacgacctggtggccgtgcacgag1920
cgcagccacctgatccagatgcccggcgagaccgagaccaagcagctgcgcaagctgcgc1980
gaggagcgccaggccgtgctgcgccccctgttcgcccagctggccctgctgcgcctgctg2040
gtgcgctgcggcgccgccgacgagcgcatccgcacccgcagctggcagcgcctgaccaag2100
cagggccgcgagttcaccaagcgcctgacccccagctggcgcgaggccctggagctggag2160
ctgacccgcctggaggcctactgcggccgcgtgcccgacgacgagtggagccgcatcgtg2220
gaccgcaccgtgatcgccctgtggcgccgcatgggcaagcaggtgcgcgactggcgcaag2280
caggtgaagagcggcgccaaggtgaaggtgaagggctaccagctggacgtggtgggcggc2340
aacagcctggcccagatcgactacctggagcagcagtacaagttcctgcgccgctggagc2400
ttcttcgcccgcgccagcggcctggtggtgcgcgccgaccgcgagagccacttcgccgtg2460
gccctgcgccagcacatcgagaacgccaagcgcgaccgcctgaagaagctggccgaccgc2520
atcctgatggaggccctgggctacgtgtacgaggccagcggcccccgcgagggccagtgg2580
accgcccagcaccccccctgccagctgatcatcctggaggagctgagcgcctaccgcttc2640
agcgacgaccgcccccccagcgagaacagcaagctgatggcctggggccaccgcggcatc2700
ctggaggagctggtgaaccaggcccaggtgcacgacgtgctggtgggcaccgtgtacgcc2760
gccttcagcagccgcttcgacgcccgcaccggcgcccccggcgtgcgctgccgccgcgtg2820
cccgcccgcttcgtgggcgccaccgtggacgacagcctgcccctgtggctgaccgagttc2880
ctggacaagcaccgcctggacaagaacctgctgcgccccgacgacgtgatccccaccggc2940
gagggcgagttcctggtgagcccctgcggcgaggaggccgcccgcgtgcgccaggtgcac3000
gccgacatcaacgccgcccagaacctgcagcgccgcctgtggcagaacttcgacatcacc3060
gagctgcgcctgcgctgcgacgtgaagatgggcggcgagggcaccgtgctggtgccccgc3120
gtgaacaacgcccgcgccaagcagctgttcggcaagaaggtgctggtgagccaggacggc3180
gtgaccttcttcgagcgcagccagaccggcggcaagccccacagcgagaagcagaccgac3240
ctgaccgacaaggagctggagctgatcgccgaggccgacgaggcccgcgccaagagcgtg3300
gtgctgttccgcgaccccagcggccacatcggcaagggccactggatccgccagcgcgag3360
ttctggagcctggtgaagcagcgcatcgagagccacaccgccgagcgcatccgcgtgcgc3420
ggcgtgggcagcagcctggac3441
<210>13
<211>3438
<212>dna
<213>alicyclobacillusmacrosporangiidus
<400>13
atgaacgtggccgtgaagagcatcaaggtgaagctgatgctgggccacctgcccgagatc60
cgcgagggcctgtggcacctgcacgaggccgtgaacctgggcgtgcgctactacaccgag120
tggctggccctgctgcgccagggcaacctgtaccgccgcggcaaggacggcgcccaggag180
tgctacatgaccgccgagcagtgccgccaggagctgctggtgcgcctgcgcgaccgccag240
aagcgcaacggccacaccggcgaccccggcaccgacgaggagctgctgggcgtggcccgc300
cgcctgtacgagctgctggtgccccagagcgtgggcaagaagggccaggcccagatgctg360
gccagcggcttcctgagccccctggccgaccccaagagcgagggcggcaagggcaccagc420
aagagcggccgcaagcccgcctggatgggcatgaaggaggccggcgacagccgctgggtg480
gaggccaaggcccgctacgaggccaacaaggccaaggaccccaccaagcaggtgatcgcc540
agcctggagatgtacggcctgcgccccctgttcgacgtgttcaccgagacctacaagacc600
atccgctggatgcccctgggcaagcaccagggcgtgcgcgcctgggaccgcgacatgttc660
cagcagagcctggagcgcctgatgagctgggagagctggaacgagcgcgtgggcgccgag720
ttcgcccgcctggtggaccgccgcgaccgcttccgcgagaagcacttcaccggccaggag780
cacctggtggccctggcccagcgcctggagcaggagatgaaggaggccagccccggcttc840
gagagcaagagcagccaggcccaccgcatcaccaagcgcgccctgcgcggcgccgacggc900
atcatcgacgactggctgaagctgagcgagggcgagcccgtggaccgcttcgacgagatc960
ctgcgcaagcgccaggcccagaacccccgccgcttcggcagccacgacctgttcctgaag1020
ctggccgagcccgtgttccagcccctgtggcgcgaggaccccagcttcctgagccgctgg1080
gccagctacaacgaggtgctgaacaagctggaggacgccaagcagttcgccaccttcacc1140
ctgcccagcccctgcagcaaccccgtgtgggcccgcttcgagaacgccgagggcaccaac1200
atcttcaagtacgacttcctgttcgaccacttcggcaagggccgccacggcgtgcgcttc1260
cagcgcatgatcgtgatgcgcgacggcgtgcccaccgaggtggagggcatcgtggtgccc1320
atcgcccccagccgccagctggacgccctggcccccaacgacgccgccagccccatcgac1380
gtgttcgtgggcgaccccgccgcccccggcgccttccgcggccagttcggcggcgccaag1440
atccagtaccgccgcagcgccctggtgcgcaagggccgccgcgaggagaaggcctacctg1500
tgcggcttccgcctgcccagccagcgccgcaccggcacccccgccgacgacgccggcgag1560
gtgttcctgaacctgagcctgcgcgtggagagccagagcgagcaggccggccgccgcaac1620
cccccctacgccgccgtgttccacatcagcgaccagacccgccgcgtgatcgtgcgctac1680
ggcgagatcgagcgctacctggccgagcaccccgacaccggcatccccggcagccgcggc1740
ctgaccagcggcctgcgcgtgatgagcgtggacctgggcctgcgcaccagcgccgccatc1800
agcgtgttccgcgtggcccaccgcgacgagctgacccccgacgcccacggccgccagccc1860
ttcttcttccccatccacggcatggaccacctggtggccctgcacgagcgcagccacctg1920
atccgcctgcccggcgagaccgagagcaagaaggtgcgcagcatccgcgagcagcgcctg1980
gaccgcctgaaccgcctgcgcagccagatggccagcctgcgcctgctggtgcgcaccggc2040
gtgctggacgagcagaagcgcgaccgcaactgggagcgcctgcagagcagcatggagcgc2100
ggcggcgagcgcatgcccagcgactggtgggacctgttccaggcccaggtgcgctacctg2160
gcccagcaccgcgacgccagcggcgaggcctggggccgcatggtgcaggccgccgtgcgc2220
accctgtggcgccagctggccaagcaggtgcgcgactggcgcaaggaggtgcgccgcaac2280
gccgacaaggtgaagatccgcggcatcgcccgcgacgtgcccggcggccacagcctggcc2340
cagctggactacctggagcgccagtaccgcttcctgcgcagctggagcgccttcagcgtg2400
caggccggccaggtggtgcgcgccgagcgcgacagccgcttcgccgtggccctgcgcgag2460
cacatcgacaacggcaagaaggaccgcctgaagaagctggccgaccgcatcctgatggag2520
gccctgggctacgtgtacgtgaccgacggccgccgcgccggccagtggcaggccgtgtac2580
cccccctgccagctggtgctgctggaggagctgagcgagtaccgcttcagcaacgaccgc2640
ccccccagcgagaacagccagctgatggtgtggagccaccgcggcgtgctggaggagctg2700
atccaccaggcccaggtgcacgacgtgctggtgggcaccatccccgccgccttcagcagc2760
cgcttcgacgcccgcaccggcgcccccggcatccgctgccgccgcgtgcccagcatcccc2820
ctgaaggacgcccccagcatccccatctggctgagccactacctgaagcagaccgagcgc2880
gacgccgccgccctgcgccccggcgagctgatccccaccggcgacggcgagttcctggtg2940
acccccgccggccgcggcgccagcggcgtgcgcgtggtgcacgccgacatcaacgccgcc3000
cacaacctgcagcgccgcctgtgggagaacttcgacctgagcgacatccgcgtgcgctgc3060
gaccgccgcgagggcaaggacggcaccgtggtgctgatcccccgcctgaccaaccagcgc3120
gtgaaggagcgctacagcggcgtgatcttcaccagcgaggacggcgtgagcttcaccgtg3180
ggcgacgccaagacccgccgccgcagcagcgccagccagggcgagggcgacgacctgagc3240
gacgaggagcaggagctgctggccgaggccgacgacgcccgcgagcgcagcgtggtgctg3300
ttccgcgaccccagcggcttcgtgaacggcggccgctggaccgcccagcgcgccttctgg3360
ggcatggtgcacaaccgcatcgagaccctgctggccgagcgcttcagcgtgagcggcgcc3420
gccgagaaggtgcgcggc3438
<210>14
<211>3324
<212>dna
<213>bacillushisashii
<400>14
atggccacccgcagcttcatcctgaagatcgagcccaacgaggaggtgaagaagggcctg60
tggaagacccacgaggtgctgaaccacggcatcgcctactacatgaacatcctgaagctg120
atccgccaggaggccatctacgagcaccacgagcaggaccccaagaaccccaagaaggtg180
agcaaggccgagatccaggccgagctgtgggacttcgtgctgaagatgcagaagtgcaac240
agcttcacccacgaggtggacaaggacgaggtgttcaacatcctgcgcgagctgtacgag300
gagctggtgcccagcagcgtggagaagaagggcgaggccaaccagctgagcaacaagttc360
ctgtaccccctggtggaccccaacagccagagcggcaagggcaccgccagcagcggccgc420
aagccccgctggtacaacctgaagatcgccggcgaccccagctgggaggaggagaagaag480
aagtgggaggaggacaagaagaaggaccccctggccaagatcctgggcaagctggccgag540
tacggcctgatccccctgttcatcccctacaccgacagcaacgagcccatcgtgaaggag600
atcaagtggatggagaagagccgcaaccagagcgtgcgccgcctggacaaggacatgttc660
atccaggccctggagcgcttcctgagctgggagagctggaacctgaaggtgaaggaggag720
tacgagaaggtggagaaggagtacaagaccctggaggagcgcatcaaggaggacatccag780
gccctgaaggccctggagcagtacgagaaggagcgccaggagcagctgctgcgcgacacc840
ctgaacaccaacgagtaccgcctgagcaagcgcggcctgcgcggctggcgcgagatcatc900
cagaagtggctgaagatggacgagaacgagcccagcgagaagtacctggaggtgttcaag960
gactaccagcgcaagcacccccgcgaggccggcgactacagcgtgtacgagttcctgagc1020
aagaaggagaaccacttcatctggcgcaaccaccccgagtacccctacctgtacgccacc1080
ttctgcgagatcgacaagaagaagaaggacgccaagcagcaggccaccttcaccctggcc1140
gaccccatcaaccaccccctgtgggtgcgcttcgaggagcgcagcggcagcaacctgaac1200
aagtaccgcatcctgaccgagcagctgcacaccgagaagctgaagaagaagctgaccgtg1260
cagctggaccgcctgatctaccccaccgagagcggcggctgggaggagaagggcaaggtg1320
gacatcgtgctgctgcccagccgccagttctacaaccagatcttcctggacatcgaggag1380
aagggcaagcacgccttcacctacaaggacgagagcatcaagttccccctgaagggcacc1440
ctgggcggcgcccgcgtgcagttcgaccgcgaccacctgcgccgctacccccacaaggtg1500
gagagcggcaacgtgggccgcatctacttcaacatgaccgtgaacatcgagcccaccgag1560
agccccgtgagcaagagcctgaagatccaccgcgacgacttccccaaggtggtgaacttc1620
aagcccaaggagctgaccgagtggatcaaggacagcaagggcaagaagctgaagagcggc1680
atcgagagcctggagatcggcctgcgcgtgatgagcatcgacctgggccagcgccaggcc1740
gccgccgccagcatcttcgaggtggtggaccagaagcccgacatcgagggcaagctgttc1800
ttccccatcaagggcaccgagctgtacgccgtgcaccgcgccagcttcaacatcaagctg1860
cccggcgagaccctggtgaagagccgcgaggtgctgcgcaaggcccgcgaggacaacctg1920
aagctgatgaaccagaagctgaacttcctgcgcaacgtgctgcacttccagcagttcgag1980
gacatcaccgagcgcgagaagcgcgtgaccaagtggatcagccgccaggagaacagcgac2040
gtgcccctggtgtaccaggacgagctgatccagatccgcgagctgatgtacaagccctac2100
aaggactgggtggccttcctgaagcagctgcacaagcgcctggaggtggagatcggcaag2160
gaggtgaagcactggcgcaagagcctgagcgacggccgcaagggcctgtacggcatcagc2220
ctgaagaacatcgacgagatcgaccgcacccgcaagttcctgctgcgctggagcctgcgc2280
cccaccgagcccggcgaggtgcgccgcctggagcccggccagcgcttcgccatcgaccag2340
ctgaaccacctgaacgccctgaaggaggaccgcctgaagaagatggccaacaccatcatc2400
atgcacgccctgggctactgctacgacgtgcgcaagaagaagtggcaggccaagaacccc2460
gcctgccagatcatcctgttcgaggacctgagcaactacaacccctacgaggagcgcagc2520
cgcttcgagaacagcaagctgatgaagtggagccgccgcgagatcccccgccaggtggcc2580
ctgcagggcgagatctacggcctgcaggtgggcgaggtgggcgcccagttcagcagccgc2640
ttccacgccaagaccggcagccccggcatccgctgcagcgtggtgaccaaggagaagctg2700
caggacaaccgcttcttcaagaacctgcagcgcgagggccgcctgaccctggacaagatc2760
gccgtgctgaaggagggcgacctgtaccccgacaagggcggcgagaagttcatcagcctg2820
agcaaggaccgcaagtgcgtgaccacccacgccgacatcaacgccgcccagaacctgcag2880
aagcgcttctggacccgcacccacggcttctacaaggtgtactgcaaggcctaccaggtg2940
gacggccagaccgtgtacatccccgagagcaaggaccagaagcagaagatcatcgaggag3000
ttcggcgagggctacttcatcctgaaggacggcgtgtacgagtgggtgaacgccggcaag3060
ctgaagatcaagaagggcagcagcaagcagagcagcagcgagctggtggacagcgacatc3120
ctgaaggacagcttcgacctggccagcgagctgaagggcgagaagctgatgctgtaccgc3180
gaccccagcggcaacgtgttccccagcgacaagtggatggccgccggcgtgttcttcggc3240
aagctggagcgcatcctgatcagcaagctgaccaaccagtacagcatcagcaccatcgag3300
gacgacagcagcaagcagagcatg3324
<210>15
<211>3324
<212>dna
<213>bacillus
<400>15
atggccatccgcagcatcaagctgaagctgaagacccacaccggccccgaggcccagaac60
ctgcgcaagggcatctggcgcacccaccgcctgctgaacgagggcgtggcctactacatg120
aagatgctgctgctgttccgccaggagagcaccggcgagcgccccaaggaggagctgcag180
gaggagctgatctgccacatccgcgagcagcagcagcgcaaccaggccgacaagaacacc240
caggccctgcccctggacaaggccctggaggccctgcgccagctgtacgagctgctggtg300
cccagcagcgtgggccagagcggcgacgcccagatcatcagccgcaagttcctgagcccc360
ctggtggaccccaacagcgagggcggcaagggcaccagcaaggccggcgccaagcccacc420
tggcagaagaagaaggaggccaacgaccccacctgggagcaggactacgagaagtggaag480
aagcgccgcgaggaggaccccaccgccagcgtgatcaccaccctggaggagtacggcatc540
cgccccatcttccccctgtacaccaacaccgtgaccgacatcgcctggctgcccctgcag600
agcaaccagttcgtgcgcacctgggaccgcgacatgctgcagcaggccatcgagcgcctg660
ctgagctgggagagctggaacaagcgcgtgcaggaggagtacgccaagctgaaggagaag720
atggcccagctgaacgagcagctggagggcggccaggagtggatcagcctgctggagcag780
tacgaggagaaccgcgagcgcgagctgcgcgagaacatgaccgccgccaacgacaagtac840
cgcatcaccaagcgccagatgaagggctggaacgagctgtacgagctgtggagcaccttc900
cccgccagcgccagccacgagcagtacaaggaggccctgaagcgcgtgcagcagcgcctg960
cgcggccgcttcggcgacgcccacttcttccagtacctgatggaggagaagaaccgcctg1020
atctggaagggcaacccccagcgcatccactacttcgtggcccgcaacgagctgaccaag1080
cgcctggaggaggccaagcagagcgccaccatgaccctgcccaacgcccgcaagcacccc1140
ctgtgggtgcgcttcgacgcccgcggcggcaacctgcaggactactacctgaccgccgag1200
gccgacaagccccgcagccgccgcttcgtgaccttcagccagctgatctggcccagcgag1260
agcggctggatggagaagaaggacgtggaggtggagctggccctgagccgccagttctac1320
cagcaggtgaagctgctgaagaacgacaagggcaagcagaagatcgagttcaaggacaag1380
ggcagcggcagcaccttcaacggccacctgggcggcgccaagctgcagctggagcgcggc1440
gacctggagaaggaggagaagaacttcgaggacggcgagatcggcagcgtgtacctgaac1500
gtggtgatcgacttcgagcccctgcaggaggtgaagaacggccgcgtgcaggccccctac1560
ggccaggtgctgcagctgatccgccgccccaacgagttccccaaggtgaccacctacaag1620
agcgagcagctggtggagtggatcaaggccagcccccagcacagcgccggcgtggagagc1680
ctggccagcggcttccgcgtgatgagcatcgacctgggcctgcgcgccgccgccgccacc1740
agcatcttcagcgtggaggagagcagcgacaagaacgccgccgacttcagctactggatc1800
gagggcacccccctggtggccgtgcaccagcgcagctacatgctgcgcctgcccggcgag1860
caggtggagaagcaggtgatggagaagcgcgacgagcgcttccagctgcaccagcgcgtg1920
aagttccagatccgcgtgctggcccagatcatgcgcatggccaacaagcagtacggcgac1980
cgctgggacgagctggacagcctgaagcaggccgtggagcagaagaagagccccctggac2040
cagaccgaccgcaccttctgggagggcatcgtgtgcgacctgaccaaggtgctgccccgc2100
aacgaggccgactgggagcaggccgtggtgcagatccaccgcaaggccgaggagtacgtg2160
ggcaaggccgtgcaggcctggcgcaagcgcttcgccgccgacgagcgcaagggcatcgcc2220
ggcctgagcatgtggaacatcgaggagctggagggcctgcgcaagctgctgatcagctgg2280
agccgccgcacccgcaacccccaggaggtgaaccgcttcgagcgcggccacaccagccac2340
cagcgcctgctgacccacatccagaacgtgaaggaggaccgcctgaagcagctgagccac2400
gccatcgtgatgaccgccctgggctacgtgtacgacgagcgcaagcaggagtggtgcgcc2460
gagtaccccgcctgccaggtgatcctgttcgagaacctgagccagtaccgcagcaacctg2520
gaccgcagcaccaaggagaacagcaccctgatgaagtgggcccaccgcagcatccccaag2580
tacgtgcacatgcaggccgagccctacggcatccagatcggcgacgtgcgcgccgagtac2640
agcagccgcttctacgccaagaccggcacccccggcatccgctgcaagaaggtgcgcggc2700
caggacctgcagggccgccgcttcgagaacctgcagaagcgcctggtgaacgagcagttc2760
ctgaccgaggagcaggtgaagcagctgcgccccggcgacatcgtgcccgacgacagcggc2820
gagctgttcatgaccctgaccgacggcagcggcagcaaggaggtggtgttcctgcaggcc2880
gacatcaacgccgcccacaacctgcagaagcgcttctggcagcgctacaacgagctgttc2940
aaggtgagctgccgcgtgatcgtgcgcgacgaggaggagtacctggtgcccaagaccaag3000
agcgtgcaggccaagctgggcaagggcctgttcgtgaagaagagcgacaccgcctggaag3060
gacgtgtacgtgtgggacagccaggccaagctgaagggcaagaccaccttcaccgaggag3120
agcgagagccccgagcagctggaggacttccaggagatcatcgaggaggccgaggaggcc3180
aagggcacctaccgcaccctgttccgcgaccccagcggcgtgttcttccccgagagcgtg3240
tggtacccccagaaggacttctggggcgaggtgaagcgcaagctgtacggcaagctgcgc3300
gagcgcttcctgaccaaggcccgc3324
<210>16
<211>3336
<212>dna
<213>bacillus
<400>16
atggccatccgcagcatcaagctgaagatgaagaccaacagcggcaccgacagcatctac60
ctgcgcaaggccctgtggcgcacccaccagctgatcaacgagggcatcgcctactacatg120
aacctgctgaccctgtaccgccaggaggccatcggcgacaagaccaaggaggcctaccag180
gccgagctgatcaacatcatccgcaaccagcagcgcaacaacggcagcagcgaggagcac240
ggcagcgaccaggagatcctggccctgctgcgccagctgtacgagctgatcatccccagc300
agcatcggcgagagcggcgacgccaaccagctgggcaacaagttcctgtaccccctggtg360
gaccccaacagccagagcggcaagggcaccagcaacgccggccgcaagccccgctggaag420
cgcctgaaggaggagggcaaccccgactgggagctggagaagaagaaggacgaggagcgc480
aaggccaaggaccccaccgtgaagatcttcgacaacctgaacaagtacggcctgctgccc540
ctgttccccctgttcaccaacatccagaaggacatcgagtggctgcccctgggcaagcgc600
cagagcgtgcgcaagtgggacaaggacatgttcatccaggccatcgagcgcctgctgagc660
tgggagagctggaaccgccgcgtggccgacgagtacaagcagctgaaggagaagaccgag720
agctactacaaggagcacctgaccggcggcgaggagtggatcgagaagatccgcaagttc780
gagaaggagcgcaacatggagctggagaagaacgccttcgcccccaacgacggctacttc840
atcaccagccgccagatccgcggctgggaccgcgtgtacgagaagtggagcaagctgccc900
gagagcgccagccccgaggagctgtggaaggtggtggccgagcagcagaacaagatgagc960
gagggcttcggcgaccccaaggtgttcagcttcctggccaaccgcgagaaccgcgacatc1020
tggcgcggccacagcgagcgcatctaccacatcgccgcctacaacggcctgcagaagaag1080
ctgagccgcaccaaggagcaggccaccttcaccctgcccgacgccatcgagcaccccctg1140
tggatccgctacgagagccccggcggcaccaacctgaacctgttcaagctggaggagaag1200
cagaagaagaactactacgtgaccctgagcaagatcatctggcccagcgaggagaagtgg1260
atcgagaaggagaacatcgagatccccctggcccccagcatccagttcaaccgccagatc1320
aagctgaagcagcacgtgaagggcaagcaggagatcagcttcagcgactacagcagccgc1380
atcagcctggacggcgtgctgggcggcagccgcatccagttcaaccgcaagtacatcaag1440
aaccacaaggagctgctgggcgagggcgacatcggccccgtgttcttcaacctggtggtg1500
gacgtggcccccctgcaggagacccgcaacggccgcctgcagagccccatcggcaaggcc1560
ctgaaggtgatcagcagcgacttcagcaaggtgatcgactacaagcccaaggagctgatg1620
gactggatgaacaccggcagcgccagcaacagcttcggcgtggccagcctgctggagggc1680
atgcgcgtgatgagcatcgacatgggccagcgcaccagcgccagcgtgagcatcttcgag1740
gtggtgaaggagctgcccaaggaccaggagcagaagctgttctacagcatcaacgacacc1800
gagctgttcgccatccacaagcgcagcttcctgctgaacctgcccggcgaggtggtgacc1860
aagaacaacaagcagcagcgccaggagcgccgcaagaagcgccagttcgtgcgcagccag1920
atccgcatgctggccaacgtgctgcgcctggagaccaagaagacccccgacgagcgcaag1980
aaggccatccacaagctgatggagatcgtgcagagctacgacagctggaccgccagccag2040
aaggaggtgtgggagaaggagctgaacctgctgaccaacatggccgccttcaacgacgag2100
atctggaaggagagcctggtggagctgcaccaccgcatcgagccctacgtgggccagatc2160
gtgagcaagtggcgcaagggcctgagcgagggccgcaagaacctggccggcatcagcatg2220
tggaacatcgacgagctggaggacacccgccgcctgctgatcagctggagcaagcgcagc2280
cgcacccccggcgaggccaaccgcatcgagaccgacgagcccttcggcagcagcctgctg2340
cagcacatccagaacgtgaaggacgaccgcctgaagcagatggccaacctgatcatcatg2400
accgccctgggcttcaagtacgacaaggaggagaaggaccgctacaagcgctggaaggag2460
acctaccccgcctgccagatcatcctgttcgagaacctgaaccgctacctgttcaacctg2520
gaccgcagccgccgcgagaacagccgcctgatgaagtgggcccaccgcagcatcccccgc2580
accgtgagcatgcagggcgagatgttcggcctgcaggtgggcgacgtgcgcagcgagtac2640
agcagccgcttccacgccaagaccggcgcccccggcatccgctgccacgccctgaccgag2700
gaggacctgaaggccggcagcaacaccctgaagcgcctgatcgaggacggcttcatcaac2760
gagagcgagctggcctacctgaagaagggcgacatcatccccagccagggcggcgagctg2820
ttcgtgaccctgagcaagcgctacaagaaggacagcgacaacaacgagctgaccgtgatc2880
cacgccgacatcaacgccgcccagaacctgcagaagcgcttctggcagcagaacagcgag2940
gtgtaccgcgtgccctgccagctggcccgcatgggcgaggacaagctgtacatccccaag3000
agccagaccgagaccatcaagaagtacttcggcaagggcagcttcgtgaagaacaacacc3060
gagcaggaggtgtacaagtgggagaagagcgagaagatgaagatcaagaccgacaccacc3120
ttcgacctgcaggacctggacggcttcgaggacatcagcaagaccatcgagctggcccag3180
gagcagcagaagaagtacctgaccatgttccgcgaccccagcggctacttcttcaacaac3240
gagacctggcgcccccagaaggagtactggagcatcgtgaacaacatcatcaagagctgc3300
ctgaagaagaagatcctgagcaacaaggtggagctg3336
<210>17
<211>3447
<212>dna
<213>desulfovibrioinopinatus
<400>17
atgcccacccgcaccatcaacctgaagctggtgctgggcaagaaccccgagaacgccacc60
ctgcgccgcgccctgttcagcacccaccgcctggtgaaccaggccaccaagcgcatcgag120
gagttcctgctgctgtgccgcggcgaggcctaccgcaccgtggacaacgagggcaaggag180
gccgagatcccccgccacgccgtgcaggaggaggccctggccttcgccaaggccgcccag240
cgccacaacggctgcatcagcacctacgaggaccaggagatcctggacgtgctgcgccag300
ctgtacgagcgcctggtgcccagcgtgaacgagaacaacgaggccggcgacgcccaggcc360
gccaacgcctgggtgagccccctgatgagcgccgagagcgagggcggcctgagcgtgtac420
gacaaggtgctggaccccccccccgtgtggatgaagctgaaggaggagaaggcccccggc480
tgggaggccgccagccagatctggatccagagcgacgagggccagagcctgctgaacaag540
cccggcagccccccccgctggatccgcaagctgcgcagcggccagccctggcaggacgac600
ttcgtgagcgaccagaagaagaagcaggacgagctgaccaagggcaacgcccccctgatc660
aagcagctgaaggagatgggcctgctgcccctggtgaaccccttcttccgccacctgctg720
gaccccgagggcaagggcgtgagcccctgggaccgcctggccgtgcgcgccgccgtggcc780
cacttcatcagctgggagagctggaaccaccgcacccgcgccgagtacaacagcctgaag840
ctgcgccgcgacgagttcgaggccgccagcgacgagttcaaggacgacttcaccctgctg900
cgccagtacgaggccaagcgccacagcaccctgaagagcatcgccctggccgacgacagc960
aacccctaccgcatcggcgtgcgcagcctgcgcgcctggaaccgcgtgcgcgaggagtgg1020
atcgacaagggcgccaccgaggagcagcgcgtgaccatcctgagcaagctgcagacccag1080
ctgcgcggcaagttcggcgaccccgacctgttcaactggctggcccaggaccgccacgtg1140
cacctgtggagcccccgcgacagcgtgacccccctggtgcgcatcaacgccgtggacaag1200
gtgctgcgccgccgcaagccctacgccctgatgaccttcgcccacccccgcttccacccc1260
cgctggatcctgtacgaggcccccggcggcagcaacctgcgccagtacgccctggactgc1320
accgagaacgccctgcacatcaccctgcccctgctggtggacgacgcccacggcacctgg1380
atcgagaagaagatccgcgtgcccctggcccccagcggccagatccaggacctgaccctg1440
gagaagctggagaagaagaagaaccgcctgtactaccgcagcggcttccagcagttcgcc1500
ggcctggccggcggcgccgaggtgctgttccaccgcccctacatggagcacgacgagcgc1560
agcgaggagagcctgctggagcgccccggcgccgtgtggttcaagctgaccctggacgtg1620
gccacccaggccccccccaactggctggacggcaagggccgcgtgcgcaccccccccgag1680
gtgcaccacttcaagaccgccctgagcaacaagagcaagcacacccgcaccctgcagccc1740
ggcctgcgcgtgctgagcgtggacctgggcatgcgcaccttcgccagctgcagcgtgttc1800
gagctgatcgagggcaagcccgagaccggccgcgccttccccgtggccgacgagcgcagc1860
atggacagccccaacaagctgtgggccaagcacgagcgcagcttcaagctgaccctgccc1920
ggcgagacccccagccgcaaggaggaggaggagcgcagcatcgcccgcgccgagatctac1980
gccctgaagcgcgacatccagcgcctgaagagcctgctgcgcctgggcgaggaggacaac2040
gacaaccgccgcgacgccctgctggagcagttcttcaagggctggggcgaggaggacgtg2100
gtgcccggccaggccttcccccgcagcctgttccagggcctgggcgccgcccccttccgc2160
agcacccccgagctgtggcgccagcactgccagacctactacgacaaggccgaggcctgc2220
ctggccaagcacatcagcgactggcgcaagcgcacccgcccccgccccaccagccgcgag2280
atgtggtacaagacccgcagctaccacggcggcaagagcatctggatgctggagtacctg2340
gacgccgtgcgcaagctgctgctgagctggagcctgcgcggccgcacctacggcgccatc2400
aaccgccaggacaccgcccgcttcggcagcctggccagccgcctgctgcaccacatcaac2460
agcctgaaggaggaccgcatcaagaccggcgccgacagcatcgtgcaggccgcccgcggc2520
tacatccccctgccccacggcaagggctgggagcagcgctacgagccctgccagctgatc2580
ctgttcgaggacctggcccgctaccgcttccgcgtggaccgcccccgccgcgagaacagc2640
cagctgatgcagtggaaccaccgcgccatcgtggccgagaccaccatgcaggccgagctg2700
tacggccagatcgtggagaacaccgccgccggcttcagcagccgcttccacgccgccacc2760
ggcgcccccggcgtgcgctgccgcttcctgctggagcgcgacttcgacaacgacctgccc2820
aagccctacctgctgcgcgagctgagctggatgctgggcaacaccaaggtggagagcgag2880
gaggagaagctgcgcctgctgagcgagaagatccgccccggcagcctggtgccctgggac2940
ggcggcgagcagttcgccaccctgcaccccaagcgccagaccctgtgcgtgatccacgcc3000
gacatgaacgccgcccagaacctgcagcgccgcttcttcggccgctgcggcgaggccttc3060
cgcctggtgtgccagccccacggcgacgacgtgctgcgcctggccagcacccccggcgcc3120
cgcctgctgggcgccctgcagcagctggagaacggccagggcgccttcgagctggtgcgc3180
gacatgggcagcaccagccagatgaaccgcttcgtgatgaagagcctgggcaagaagaag3240
atcaagcccctgcaggacaacaacggcgacgacgagctggaggacgtgctgagcgtgctg3300
cccgaggaggacgacaccggccgcatcaccgtgttccgcgacagcagcggcatcttcttc3360
ccctgcaacgtgtggatccccgccaagcagttctggcccgccgtgcgcgccatgatctgg3420
aaggtgatggccagccacagcctgggc3447
<210>18
<211>3270
<212>dna
<213>laceyellasediminis
<400>18
atgagcatccgcagcttcaagctgaagatcaagaccaagagcggcgtgaacgccgaggag60
ctgcgccgcggcctgtggcgcacccaccagctgatcaacgacggcatcgcctactacatg120
aactggctggtgctgctgcgccaggaggacctgttcatccgcaacgaggagaccaacgag180
atcgagaagcgcagcaaggaggagatccagggcgagctgctggagcgcgtgcacaagcag240
cagcagcgcaaccagtggagcggcgaggtggacgaccagaccctgctgcagaccctgcgc300
cacctgtacgaggagatcgtgcccagcgtgatcggcaagagcggcaacgccagcctgaag360
gcccgcttcttcctgggccccctggtggaccccaacaacaagaccaccaaggacgtgagc420
aagagcggccccacccccaagtggaagaagatgaaggacgccggcgaccccaactgggtg480
caggagtacgagaagtacatggccgagcgccagaccctggtgcgcctggaggagatgggc540
ctgatccccctgttccccatgtacaccgacgaggtgggcgacatccactggctgccccag600
gccagcggctacacccgcacctgggaccgcgacatgttccagcaggccatcgagcgcctg660
ctgagctgggagagctggaaccgccgcgtgcgcgagcgccgcgcccagttcgagaagaag720
acccacgacttcgccagccgcttcagcgagagcgacgtgcagtggatgaacaagctgcgc780
gagtacgaggcccagcaggagaagagcctggaggagaacgccttcgcccccaacgagccc840
tacgccctgaccaagaaggccctgcgcggctgggagcgcgtgtaccacagctggatgcgc900
ctggacagcgccgccagcgaggaggcctactggcaggaggtggccacctgccagaccgcc960
atgcgcggcgagttcggcgaccccgccatctaccagttcctggcccagaaggagaaccac1020
gacatctggcgcggctaccccgagcgcgtgatcgacttcgccgagctgaaccacctgcag1080
cgcgagctgcgccgcgccaaggaggacgccaccttcaccctgcccgacagcgtggaccac1140
cccctgtgggtgcgctacgaggcccccggcggcaccaacatccacggctacgacctggtg1200
caggacaccaagcgcaacctgaccctgatcctggacaagttcatcctgcccgacgagaac1260
ggcagctggcacgaggtgaagaaggtgcccttcagcctggccaagagcaagcagttccac1320
cgccaggtgtggctgcaggaggagcagaagcagaagaagcgcgaggtggtgttctacgac1380
tacagcaccaacctgccccacctgggcaccctggccggcgccaagctgcagtgggaccgc1440
aacttcctgaacaagcgcacccagcagcagatcgaggagaccggcgagatcggcaaggtg1500
ttcttcaacatcagcgtggacgtgcgccccgccgtggaggtgaagaacggccgcctgcag1560
aacggcctgggcaaggccctgaccgtgctgacccaccccgacggcaccaagatcgtgacc1620
ggctggaaggccgagcagctggagaagtgggtgggcgagagcggccgcgtgagcagcctg1680
ggcctggacagcctgagcgagggcctgcgcgtgatgagcatcgacctgggccagcgcacc1740
agcgccaccgtgagcgtgttcgagatcaccaaggaggcccccgacaacccctacaagttc1800
ttctaccagctggagggcaccgagctgttcgccgtgcaccagcgcagcttcctgctggcc1860
ctgcccggcgagaaccccccccagaagatcaagcagatgcgcgagatccgctggaaggag1920
cgcaaccgcatcaagcagcaggtggaccagctgagcgccatcctgcgcctgcacaagaag1980
gtgaacgaggacgagcgcatccaggccatcgacaagctgctgcagaaggtggccagctgg2040
cagctgaacgaggagatcgccaccgcctggaaccaggccctgagccagctgtacagcaag2100
gccaaggagaacgacctgcagtggaaccaggccatcaagaacgcccaccaccagctggag2160
cccgtggtgggcaagcagatcagcctgtggcgcaaggacctgagcaccggccgccagggc2220
atcgccggcctgagcctgtggagcatcgaggagctggaggccaccaagaagctgctgacc2280
cgctggagcaagcgcagccgcgagcccggcgtggtgaagcgcatcgagcgcttcgagacc2340
ttcgccaagcagatccagcaccacatcaaccaggtgaaggagaaccgcctgaagcagctg2400
gccaacctgatcgtgatgaccgccctgggctacaagtacgaccaggagcagaagaagtgg2460
atcgaggtgtaccccgcctgccaggtggtgctgttcgagaacctgcgcagctaccgcttc2520
agctacgagcgcagccgccgcgagaacaagaagctgatggagtggagccaccgcagcatc2580
cccaagctggtgcagatgcagggcgagctgttcggcctgcaggtggccgacgtgtacgcc2640
gcctacagcagccgctaccacggccgcaccggcgcccccggcatccgctgccacgccctg2700
accgaggccgacctgcgcaacgagaccaacatcatccacgagctgatcgaggccggcttc2760
atcaaggaggagcaccgcccctacctgcagcagggcgacctggtgccctggagcggcggc2820
gagctgttcgccaccctgcagaagccctacgacaacccccgcatcctgaccctgcacgcc2880
gacatcaacgccgcccagaacatccagaagcgcttctggcaccccagcatgtggttccgc2940
gtgaactgcgagagcgtgatggagggcgagatcgtgacctacgtgcccaagaacaagacc3000
gtgcacaagaagcagggcaagaccttccgcttcgtgaaggtggagggcagcgacgtgtac3060
gagtgggccaagtggagcaagaaccgcaacaagaacaccttcagcagcatcaccgagcgc3120
aagccccccagcagcatgatcctgttccgcgaccccagcggcaccttcttcaaggagcag3180
gagtgggtggagcagaagaccttctggggcaaggtgcagagcatgatccaggcctacatg3240
aagaagaccatcgtgcagcgcatggaggag3270
<210>19
<211>3357
<212>dna
<213>spirochaetes
<400>19
atgagcttcaccatcagctaccccttcaagctgatcatcaagaacaaggacgaggccaag60
gccctgctggacacccaccagtacatgaacgagggcgtgaagtactacctggagaagctg120
ctgatgttccgccaggagaagatcttcatcggcgaggacgagaccggcaagcgcatctac180
atcgaggagaccgagtacaagaagcagatcgaggagttctacctgatcaagaagaccgag240
ctgggccgcaacctgaccctgaccctggacgagttcaagaccctgatgcgcgagctgtac300
atctgcctggtgagcagcagcatggagaacaagaagggcttccccaacgcccagcaggcc360
agcctgaacatcttcagccccctgttcgacgccgagagcaagggctacatcctgaaggag420
gagaacaacaacatcagcctgatccacaaggactacggcaagatcctgctgaagcgcctg480
cgcgacaacaacctgatccccatcttcaccaagttcaccgacatcaagaagatcaccgcc540
aagctgagccccaccgccctggaccgcatgatcttcgcccaggccatcgagaagctgctg600
agctacgagagctggtgcaagctgatgatcaaggagcgcttcgacaaggaggtgaagatc660
aaggagctggagaacaagtgcgagaacaagcaggagcgcgacaagatcttcgagatcctg720
gagaagtacgaggaggagcgccagaagaccttcgagcaggacagcggcttcgccaagaag780
ggcaagttctacatcaccggccgcatgctgaagggcttcgacgagatcaaggagaagtgg840
ctgaaggagaaggaccgcagcgagcagaacctgatcaacatcctgaacaagtaccagacc900
gacaacagcaagctggtgggcgaccgcaacctgttcgagttcatcatcaagctggagaac960
cagtgcctgtggaacggcgacatcgactacctgaagatcaagcgcgacatcaacaagaac1020
cagatctggctggaccgccccgagatgccccgcttcaccatgcccgacttcaagaagcac1080
cccctgtggtaccgctacgaggaccccagcaacagcaacttccgcaactacaagatcgag1140
gtggtgaaggacgagaactacatcaccatccccctgatcaccgagcgcaacaacgagtac1200
ttcgaggagaactacaccttcaacctggccaagctgaagaagctgagcgagaacatcacc1260
ttcatccccaagagcaagaacaaggagttcgagttcatcgacagcaacgacgaggaggag1320
gacaagaaggaccagaagaagagcaagcagtacatcaagtactgcgacaccgccaagaac1380
accagctacggcaagagcggcggcatccgcctgtacttcaaccgcaacgagctggagaac1440
tacaaggacggcaagaagatggacagctacaccgtgttcaccctgagcatccgcgactac1500
aagagcctgttcgccaaggagaagctgcagccccagatcttcaacaccgtggacaacaag1560
atcaccagcctgaagatccagaagaagttcggcaacgaggagcagaccaacttcctgagc1620
tacttcacccagaaccagatcaccaagaaggactggatggacgagaagaccttccagaac1680
gtgaaggagctgaacgagggcatccgcgtgctgagcgtggacctgggccagcgcttcttc1740
gccgccgtgagctgcttcgagatcatgagcgagatcgacaacaacaagctgttcttcaac1800
ctgaacgaccagaaccacaagatcatccgcatcaacgacaagaactactacgccaagcac1860
atctacagcaagaccatcaagctgagcggcgaggacgacgacctgtacaaggagcgcaag1920
atcaacaagaactacaagctgagctaccaggagcgcaagaacaagatcggcatcttcacc1980
cgccagatcaacaagctgaaccagctgctgaagatcatccgcaacgacgagatcgacaag2040
gagaagttcaaggagctgatcgagaccaccaagcgctacgtgaagaacacctacaacgac2100
ggcatcatcgactggaacaacgtggacaacaagatcctgagctacgagaacaaggaggac2160
gtgatcaacctgcacaaggagctggacaagaagctggagatcgacttcaaggagttcatc2220
cgcgagtgccgcaagcccatcttccgcagcggcggcctgagcatgcagcgcatcgacttc2280
ctggagaagctgaacaagctgaagcgcaagtgggtggcccgcacccagaagagcgccgag2340
agcatcgtgctgacccccaagttcggctacaagctgaaggagcacatcaacgagctgaag2400
gacaaccgcgtgaagcagggcgtgaactacatcctgatgaccgccctgggctacatcaag2460
gacaacgagatcaagaacgacagcaagaagaagcagaaggaggactgggtgaagaagaac2520
cgcgcctgccagatcatcctgatggagaagctgaccgagtacaccttcgccgaggaccgc2580
ccccgcgaggagaacagcaagctgcgcatgtggagccaccgccagatcttcaacttcctg2640
cagcagaaggccagcctgtggggcatcctggtgggcgacgtgttcgccccctacaccagc2700
aagtgcctgagcgacaacaacgcccccggcatccgctgccaccaggtgaccaagaaggac2760
ctgatcgacaacagctggttcctgaagatcgtggtgaaggacgacgccttctgcgacctg2820
atcgagatcaacaaggagaacgtgaagaacaagagcatcaagatcaacgacatcctgccc2880
ctgcgcggcggcgagctgttcgccagcatcaaggacggcaagctgcacatcgtgcaggcc2940
gacatcaacgccagccgcaacatcgccaagcgcttcctgagccagatcaaccccttccgc3000
gtggtgctgaagaaggacaaggacgagaccttccacctgaagaacgagcccaactacctg3060
aagaactactacagcatcctgaacttcgtgcccaccaacgaggagctgaccttcttcaag3120
gtggaggagaacaaggacatcaagcccaccaagcgcatcaagatggacaagcacgagaag3180
gagagcaccgacgagggcgacgactacagcaagaaccagatcgccctgttccgcgacgac3240
agcggcatcttcttcgacaagagcctgtgggtggacggcaagatcttctggagcgtggtg3300
aagaacaagatgaccaagctgctgcgcgagcgcaacaacaagaagaacggcagcaag3357
<210>20
<211>3426
<212>dna
<213>tuberibacilluscalidus
<400>20
atgaacatccacctgaaggagctgatccgcatggccaccaagagcttcatcctgaagatg60
aagaccaagaacaacccccagctgcgcctgagcctgtggaagacccacgagctgttcaac120
ttcggcgtggcctactacatggacctgctgagcctgttccgccagaaggacctgtacatg180
cacaacgacgaggaccccgaccaccccgtggtgctgaagaaggaggagatccaggagcgc240
ctgtggatgaaggtgcgcgagacccagcagaagaacggcttccacggcgaggtgagcaag300
gacgaggtgctggagaccctgcgcgccctgtacgaggagctggtgcccagcgccgtgggc360
aagagcggcgaggccaaccagatcagcaacaagtacctgtaccccctgaccgaccccgcc420
agccagagcggcaagggcaccgccaacagcggccgcaagccccgctggaagaagctgaag480
gaggccggcgaccccagctggaaggacgcctacgagaagtgggagaaggagcgccaggag540
gaccccaagctgaagatcctggccgccctgcagagcttcggcctgatccccctgttccgc600
cccttcaccgagaacgaccacaaggccgtgatcagcgtgaagtggatgcccaagagcaag660
aaccagagcgtgcgcaagttcgacaaggacatgttcaaccaggccatcgagcgcttcctg720
agctgggagagctggaacgagaaggtggccgaggactacgagaagaccgtgagcatctac780
gagagcctgcagaaggagctgaagggcatcagcaccaaggccttcgagatcatggagcgc840
gtggagaaggcctacgaggcccacctgcgcgagatcaccttcagcaacagcacctaccgc900
atcggcaaccgcgccatccgcggctggaccgagatcgtgaagaagtggatgaagctggac960
cccagcgccccccagggcaactacctggacgtggtgaaggactaccagcgccgccacccc1020
cgcgagagcggcgacttcaagctgttcgagctgctgagccgccccgagaaccaggccgcc1080
tggcgcgagtaccccgagttcctgcccctgtacgtgaagtaccgccacgccgagcagcgc1140
atgaagaccgccaagaagcaggccaccttcaccctgtgcgaccccatccgccaccccctg1200
tgggtgcgctacgaggagcgcagcggcaccaacctgaacaagtaccgcctgatcatgaac1260
gagaaggagaaggtggtgcagttcgaccgcctgatctgcctgaacgccgacggccactac1320
gaggagcaggaggacgtgaccgtgcccctggcccccagccagcagttcgacgaccagatc1380
aagttcagcagcgaggacaccggcaagggcaagcacaacttcagctactaccacaagggc1440
atcaactacgagctgaagggcaccctgggcggcgcccgcatccagttcgaccgcgagcac1500
ctgctgcgccgccagggcgtgaaggccggcaacgtgggccgcatcttcctgaacgtgacc1560
ctgaacatcgagcccatgcagcccttcagccgcagcggcaacctgcagaccagcgtgggc1620
aaggccctgaaggtgtacgtggacggctaccccaaggtggtgaacttcaagcccaaggag1680
ctgaccgagcacatcaaggagagcgagaagaacaccctgaccctgggcgtggagagcctg1740
cccaccggcctgcgcgtgatgagcgtggacctgggccagcgccaggccgccgccatcagc1800
atcttcgaggtggtgagcgagaagcccgacgacaacaagctgttctaccccgtgaaggac1860
accgacctgttcgccgtgcaccgcaccagcttcaacatcaagctgcccggcgagaagcgc1920
accgagcgccgcatgctggagcagcagaagcgcgaccaggccatccgcgacctgagccgc1980
aagctgaagttcctgaagaacgtgctgaacatgcagaagctggagaagaccgacgagcgc2040
gagaagcgcgtgaaccgctggatcaaggaccgcgagcgcgaggaggagaaccccgtgtac2100
gtgcaggagttcgagatgatcagcaaggtgctgtacagcccccacagcgtgtgggtggac2160
cagctgaagagcatccaccgcaagctggaggagcagctgggcaaggagatcagcaagtgg2220
cgccagagcatcagccagggccgccagggcgtgtacggcatcagcctgaagaacatcgag2280
gacatcgagaagacccgccgcctgctgttccgctggagcatgcgccccgagaaccccggc2340
gaggtgaagcagctgcagcccggcgagcgcttcgccatcgaccagcagaaccacctgaac2400
cacctgaaggacgaccgcatcaagaagctggccaaccagatcgtgatgaccgccctgggc2460
taccgctacgacggcaagcgcaagaagtggatcgccaagcaccccgcctgccagctggtg2520
ctgttcgaggacctgagccgctacgccttctacgacgagcgcagccgcctggagaaccgc2580
aacctgatgcgctggagccgccgcgagatccccaagcaggtggcccagatcggcggcctg2640
tacggcctgctggtgggcgaggtgggcgcccagtacagcagccgcttccacgccaagagc2700
ggcgcccccggcatccgctgccgcgtggtgaaggagcacgagctgtacatcaccgagggc2760
ggccagaaggtgcgcaaccagaagttcctggacagcctggtggagaacaacatcatcgag2820
cccgacgacgcccgccgcctggagcccggcgacctgatccgcgaccagggcggcgacaag2880
ttcgccaccctggacgagcgcggcgagctggtgatcacccacgccgacatcaacgccgcc2940
cagaacctgcagaagcgcttctggacccgcacccacggcctgtaccgcatccgctgcgag3000
agccgcgagatcaaggacgccgtggtgctggtgcccagcgacaaggaccagaaggagaag3060
atggagaacctgttcggcatcggctacctgcagcccttcaagcaggagaacgacgtgtac3120
aagtgggtgaagggcgagaagatcaagggcaagaagaccagcagccagagcgacgacaag3180
gagctggtgagcgagatcctgcaggaggcgagcgtgatggccgacgagctgaagggcaac3240
cgcaagaccctgttccgcgaccccagcggctacgtgttccccaaggaccgctggtacacc3300
ggcggccgctacttcggcaccctggagcacctgctgaagcgcaagctggccgagcgccgc3360
ctgttcgacggcggcagcagccgccgcggcctgttcaacggcaccgacagcaacaccaac3420
gtggag3426
<210>21
<211>2870
<212>dna
<213>artificialsequence
<220>
- 还没有人留言评论。精彩留言会获得点赞!